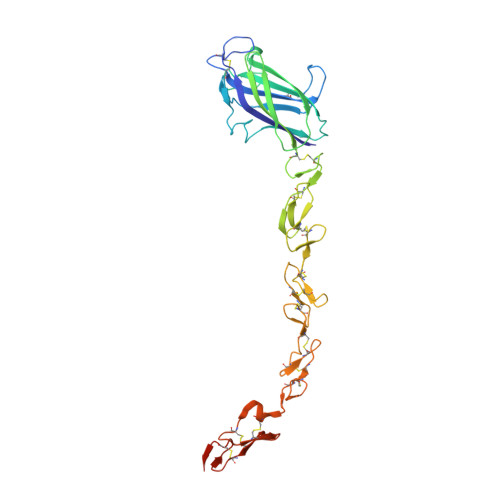
Structural and functional dissection of the interplay between lipid and Notch binding by human Notch ligands.
Suckling, R.J., Korona, B., Whiteman, P., Chillakuri, C., Holt, L., Handford, P.A., Lea, S.M.(2017) EMBO J 36: 2204-2215
- PubMed: 28572448
- DOI: https://doi.org/10.15252/embj.201796632
- Primary Citation of Related Structures:
5MVX, 5MW5, 5MW7, 5MWB, 5MWF - PubMed Abstract:
Recent data have expanded our understanding of Notch signalling by identifying a C2 domain at the N-terminus of Notch ligands, which has both lipid- and receptor-binding properties. We present novel structures of human ligands Jagged2 and Delta-like4 and human Notch2, together with functional assays, which suggest that ligand-mediated coupling of membrane recognition and Notch binding is likely to be critical in establishing the optimal context for Notch signalling. Comparisons between the Jagged and Delta family show a huge diversity in the structures of the loops at the apex of the C2 domain implicated in membrane recognition and Jagged1 missense mutations, which affect these loops and are associated with extrahepatic biliary atresia, lead to a loss of membrane recognition, but do not alter Notch binding. Taken together, these data suggest that C2 domain binding to membranes is an important element in tuning ligand-dependent Notch signalling in different physiological contexts.
Organizational Affiliation:
Sir William Dunn School of Pathology, University of Oxford, Oxford, UK.