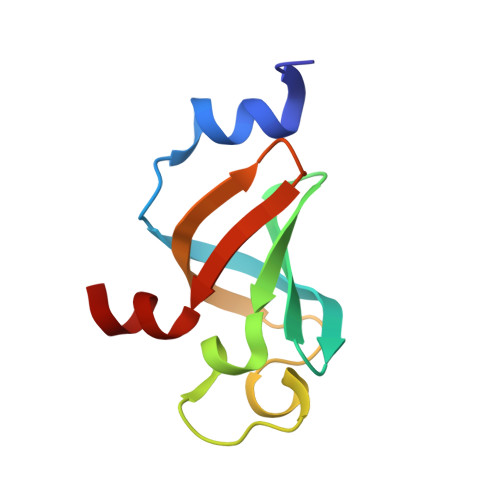
(1)H, (15)N, (13)C resonance assignments for pyrazinoic acid binding domain of ribosomal protein S1 from Mycobacterium tuberculosis
Huang, B., Fu, J., Guo, C., Wu, X., Lin, D., Liao, X.(2016) Biomol NMR Assign 10: 321-324
- PubMed: 27412769
- DOI: https://doi.org/10.1007/s12104-016-9692-9
- Primary Citation of Related Structures:
5IE8 - PubMed Abstract:
Ribosomal protein S1 of Mycobacterium tuberculosis (MtRpsA) binds to ribosome and mRNA, and plays significant role in the regulation of translation initiation, conventional protein synthesis and transfer-messenger RNA (tmRNA) mediated trans-translation. It has been identified as the target of pyrazinoic acid (POA), a bactericidal moiety from hydrolysis of pyrazinamide, which is a mainstay of combination therapy for tuberculosis. POA prevented the interactions between the C-terminal S1 domain of MtRpsA (residues 280-368, MtRpsA(CTD)_S1) and tmRNA; so that POA can inhibit the trans-translation, which is a key component of multiple quality control pathways in bacteria. However, the details of molecular mechanism and dynamic characteristics for MtRpsA(CTD)_S1 interactions with POA, tmRNA or mRNA are still unclear. Here we present the (1)H, (15)N, (13)C resonance assignments of MtRpsA(CTD)_S1 as well as the secondary structure information based on backbone chemical shifts, which lay foundation for further solution structure determination, dynamic properties characterization and interactions investigation between MtRpsA(CTD)_S1 and tmRNA, RNA or POA.
Organizational Affiliation:
Key Laboratory of Chemical Biology of Fujian Province, College of Chemistry and Chemical Engineering, Xiamen University, Xiamen, 361005, China.