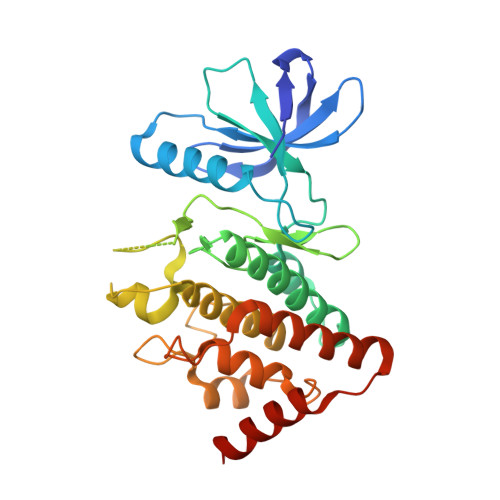
Structural Basis for Noncanonical Substrate Recognition of Cofilin/ADF Proteins by LIM Kinases.
Hamill, S., Lou, H.J., Turk, B.E., Boggon, T.J.(2016) Mol Cell 62: 397-408
- PubMed: 27153537
- DOI: https://doi.org/10.1016/j.molcel.2016.04.001
- Primary Citation of Related Structures:
5HVJ, 5HVK - PubMed Abstract:
Cofilin/actin-depolymerizing factor (ADF) proteins are critical nodes that relay signals from protein kinase cascades to the actin cytoskeleton, in particular through site-specific phosphorylation at residue Ser3. This is important for regulation of the roles of cofilin in severing and stabilizing actin filaments. Consequently, cofilin/ADF Ser3 phosphorylation is tightly controlled as an almost exclusive substrate for LIM kinases. Here we determine the LIMK1:cofilin-1 co-crystal structure. We find an interface that is distinct from canonical kinase-substrate interactions. We validate this previously unobserved mechanism for high-fidelity kinase-substrate recognition by in vitro kinase assays, examination of cofilin phosphorylation in mammalian cells, and functional analysis in S. cerevisiae. The interface is conserved across all LIM kinases. Remarkably, we also observe both pre- and postphosphotransfer states in the same crystal lattice. This study therefore provides a molecular understanding of how kinase-substrate recognition acts as a gatekeeper to regulate actin cytoskeletal dynamics.
Organizational Affiliation:
Department of Pharmacology, Yale University School of Medicine, New Haven, CT 06520, USA.