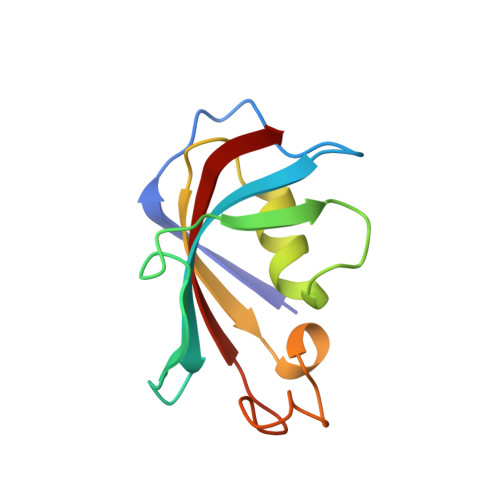
Total chemical synthesis, refolding, and crystallographic structure of fully active immunophilin calstabin 2 (FKBP12.6).
Bacchi, M., Jullian, M., Sirigu, S., Fould, B., Huet, T., Bruyand, L., Antoine, M., Vuillard, L., Ronga, L., Chavas, L.M., Nosjean, O., Ferry, G., Puget, K., Boutin, J.A.(2016) Protein Sci 25: 2225-2242
- PubMed: 27670942
- DOI: https://doi.org/10.1002/pro.3051
- Primary Citation of Related Structures:
5HKG - PubMed Abstract:
Synthetic biology (or chemical biology) is a growing field to which the chemical synthesis of proteins, particularly enzymes, makes a fundamental contribution. However, the chemical synthesis of catalytically active proteins (enzymes) remains poorly documented because it is difficult to obtain enough material for biochemical experiments. We chose calstabin, a 107-amino-acid proline isomerase, as a model. We synthesized the enzyme using the native chemical ligation approach and obtained several tens of milligrams. The polypeptide was refolded properly, and we characterized its biophysical properties, measured its catalytic activity, and then crystallized it in order to obtain its tridimensional structure after X-ray diffraction. The refolded enzyme was compared to the recombinant, wild-type enzyme. In addition, as a first step of validating the whole process, we incorporated exotic amino acids into the N-terminus. Surprisingly, none of the changes altered the catalytic activities of the corresponding mutants. Using this body of techniques, avenues are now open to further obtain enzymes modified with exotic amino acids in a way that is only barely accessible by molecular biology, obtaining detailed information on the structure-function relationship of enzymes reachable by complete chemical synthesis.
Organizational Affiliation:
Pôle d'Expertise Biotechnologie, Chimie and Biologie, Institut de Recherches Servier, 125 Chemin de Ronde, Croissy-sur-Seine, 78290, France.