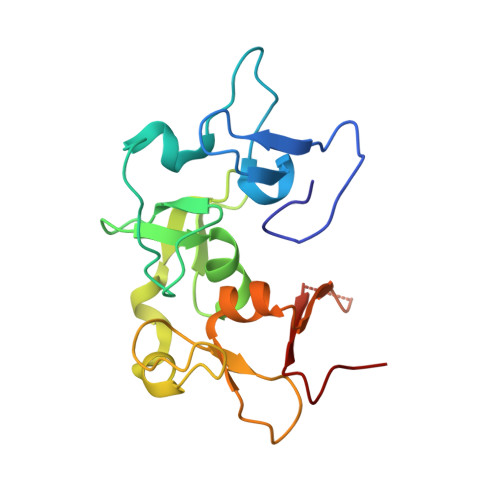
Bivalent interaction of the PZP domain of BRPF1 with the nucleosome impacts chromatin dynamics and acetylation.
Klein, B.J., Muthurajan, U.M., Lalonde, M.E., Gibson, M.D., Andrews, F.H., Hepler, M., Machida, S., Yan, K., Kurumizaka, H., Poirier, M.G., Cote, J., Luger, K., Kutateladze, T.G.(2016) Nucleic Acids Res 44: 472-484
- PubMed: 26626149
- DOI: https://doi.org/10.1093/nar/gkv1321
- Primary Citation of Related Structures:
5ERC - PubMed Abstract:
BRPF1 (bromodomain PHD finger 1) is a core subunit of the MOZ histone acetyltransferase (HAT) complex, critical for normal developmental programs and implicated in acute leukemias. BRPF1 contains a unique assembly of zinc fingers, termed a PZP domain, the physiological role of which remains unclear. Here, we elucidate the structure-function relationship of this novel epigenetic reader and detail the biological and mechanistic consequences of its interaction with nucleosomes. PZP has a globular architecture and forms a 2:1 stoichiometry complex with the nucleosome, bivalently interacting with histone H3 and DNA. This binding impacts the nucleosome dynamics, shifting the DNA unwrapping/rewrapping equilibrium toward the unwrapped state and increasing DNA accessibility. We demonstrate that the DNA-binding function of the BRPF1 PZP domain is required for the MOZ-BRPF1-ING5-hEaf6 HAT complex to be recruited to chromatin and to acetylate nucleosomal histones. Our findings reveal a novel link between chromatin dynamics and MOZ-mediated acetylation.
Organizational Affiliation:
Department of Pharmacology, University of Colorado School of Medicine, Aurora, CO 80045, USA.