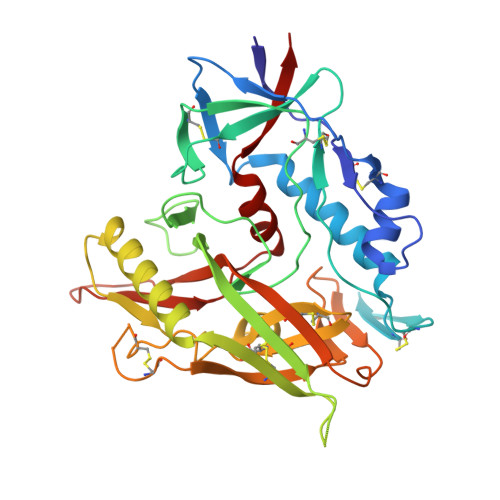
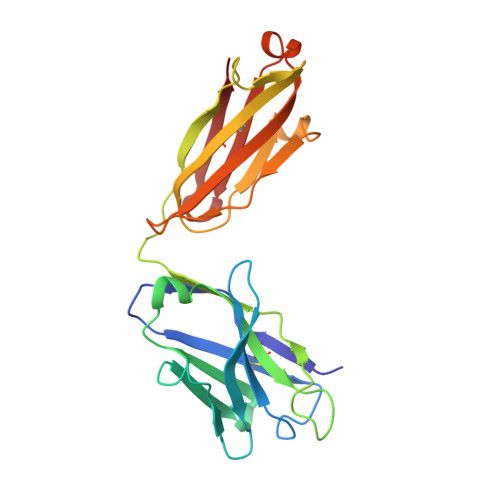
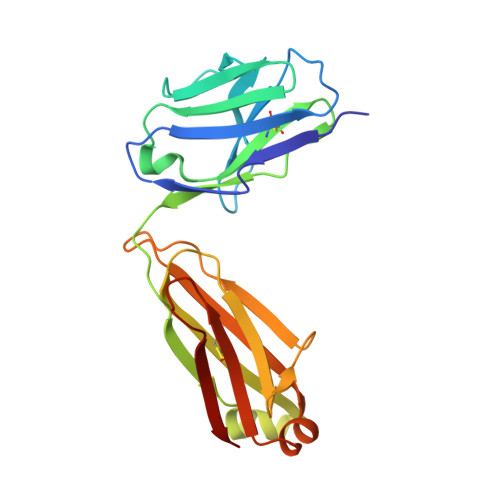
Key gp120 Glycans Pose Roadblocks to the Rapid Development of VRC01-Class Antibodies in an HIV-1-Infected Chinese Donor.
Kong, L., Ju, B., Chen, Y., He, L., Ren, L., Liu, J., Hong, K., Su, B., Wang, Z., Ozorowski, G., Ji, X., Hua, Y., Chen, Y., Deller, M.C., Hao, Y., Feng, Y., Garces, F., Wilson, R., Dai, K., O'Dell, S., McKee, K., Mascola, J.R., Ward, A.B., Wyatt, R.T., Li, Y., Wilson, I.A., Zhu, J., Shao, Y.(2016) Immunity 44: 939-950
- PubMed: 27067056
- DOI: https://doi.org/10.1016/j.immuni.2016.03.006
- Primary Citation of Related Structures:
5CD3, 5CD5 - PubMed Abstract:
VRC01-class antibodies neutralize diverse HIV-1 strains by targeting the conserved CD4-binding site. Despite extensive investigations, crucial events in the early stage of VRC01 development remain elusive. We demonstrated how VRC01-class antibodies emerged in a Chinese donor by antigen-specific single B cell sorting, structural and functional studies, and longitudinal antibody and virus repertoire analyses. A monoclonal antibody DRVIA7 with modest neutralizing breadth was isolated that displayed a subset of VRC01 signatures. X-ray and EM structures revealed a VRC01-like angle of approach, but less favorable interactions between the DRVIA7 light-chain CDR1 and the N terminus with N276 and V5 glycans of gp120. Although the DRVIA7 lineage was unable to acquire broad neutralization, longitudinal analysis revealed a repertoire-encoded VRC01 light-chain CDR3 signature and VRC01-like neutralizing heavy-chain precursors that rapidly matured within 2 years. Thus, light chain accommodation of the glycan shield should be taken into account in vaccine design targeting this conserved site of vulnerability.
Organizational Affiliation:
Department of Integrative Structural and Computational Biology, The Scripps Research Insititute, La Jolla, CA 92037, USA; Scripps Center for HIV/AIDS Vaccine Immunology & Immunogen Discovery, The Scripps Research Institute, La Jolla, CA 92037, USA; International AIDS Vaccine Initiative Neutralizing Antibody Center and the Collaboration for AIDS Vaccine Discovery, The Scripps Research Institute, La Jolla, CA 92037, USA.