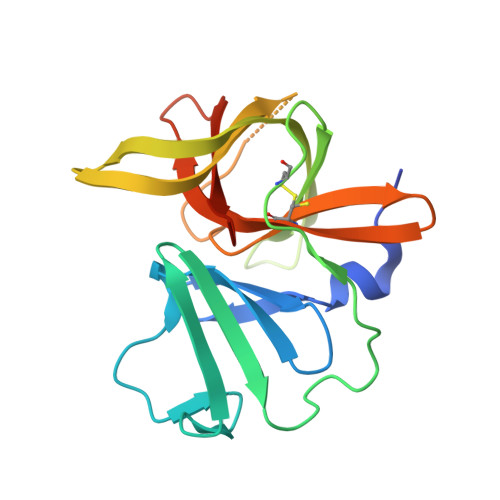
Structure-Guided Design and Optimization of Dipeptidyl Inhibitors of Norovirus 3CL Protease. Structure-Activity Relationships and Biochemical, X-ray Crystallographic, Cell-Based, and In Vivo Studies.
Galasiti Kankanamalage, A.C., Kim, Y., Weerawarna, P.M., Uy, R.A., Damalanka, V.C., Mandadapu, S.R., Alliston, K.R., Mehzabeen, N., Battaile, K.P., Lovell, S., Chang, K.O., Groutas, W.C.(2015) J Med Chem 58: 3144-3155
- PubMed: 25761614
- DOI: https://doi.org/10.1021/jm5019934
- Primary Citation of Related Structures:
4XBB, 4XBC, 4XBD - PubMed Abstract:
Norovirus infection constitutes the primary cause of acute viral gastroenteritis. There are currently no vaccines or norovirus-specific antiviral therapeutics available for the management of norovirus infection. Norovirus 3C-like protease is essential for viral replication, consequently, inhibition of this enzyme is a fruitful avenue of investigation that may lead to the emergence of antinorovirus therapeutics. We describe herein the optimization of dipeptidyl inhibitors of norovirus 3C-like protease using iterative SAR, X-ray crystallographic, and enzyme and cell-based studies. We also demonstrate herein in vivo efficacy of an inhibitor using the murine model of norovirus infection.
Organizational Affiliation:
†Department of Chemistry, Wichita State University, 1845 North Fairmount Avenue, Wichita, Kansas 67260, United States.