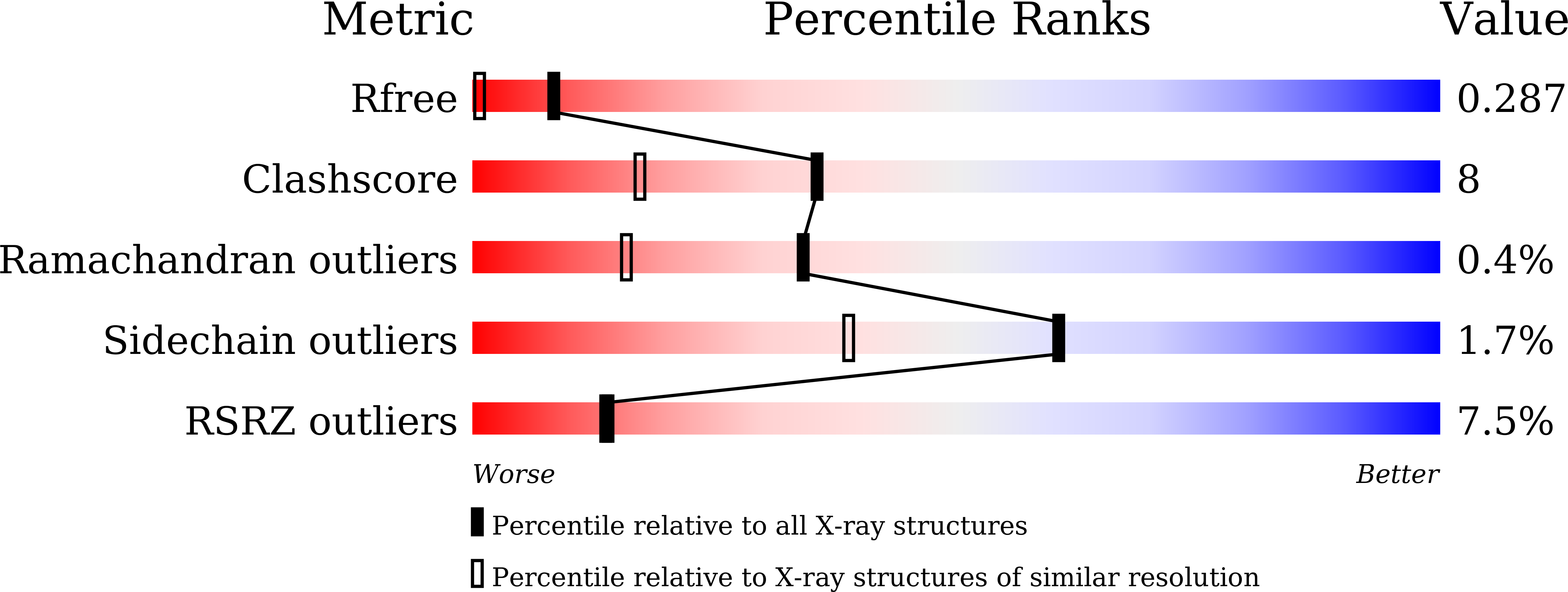
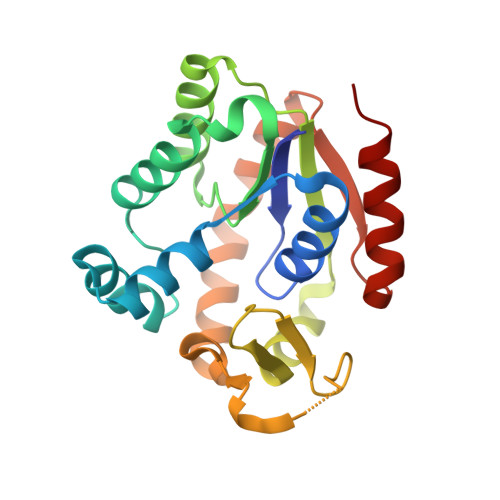
New crystal structures of adenylate kinase from Streptococcus pneumoniae D39 in two conformations.
Thach, T.T., Lee, S.(2014) Acta Crystallogr F Struct Biol Commun 70: 1468-1471
- PubMed: 25372811
- DOI: https://doi.org/10.1107/S2053230X14020718
- Primary Citation of Related Structures:
4W5H, 4W5J - PubMed Abstract:
Adenylate kinases (AdKs; EC 2.7.3.4) play a critical role in intercellular homeostasis by the interconversion of ATP and AMP to two ADP molecules. Crystal structures of adenylate kinase from Streptococcus pneumoniae D39 (SpAdK) have recently been determined using ligand-free and inhibitor-bound crystals belonging to space groups P21 and P1, respectively. Here, new crystal structures of SpAdK in ligand-free and inhibitor-bound states determined at 1.96 and 1.65 Å resolution, respectively, are reported. The new ligand-free crystal belonged to space group C2, with unit-cell parameters a=73.5, b=54.3, c=62.7 Å, β=118.8°. The new ligand-free structure revealed an open conformation that differed from the previously determined conformation, with an r.m.s.d on Cα atoms of 1.4 Å. The new crystal of the complex with the two-substrate-mimicking inhibitor P1,P5-bis(adenosine-5'-)pentaphosphate (Ap5A) belonged to space group P1, with unit-cell parameters a=53.9, b=62.3, c=63.0 Å, α=101.9, β=112.6, γ=89.9°. Despite belonging to the same space group as the previously reported crystal, the new Ap5A-bound crystal contains four molecules in the asymmetric unit, compared with two in the previous crystal, and shows slightly different lattice contacts. These results demonstrate that SpAdK can crystallize promiscuously in different forms and that the open structure is flexible in conformation.
Organizational Affiliation:
Department of Biological Sciences, Sungkyunkwan University, 2066 Seobu-ro, Suwon, Gyeonggi 440-746, Republic of Korea.