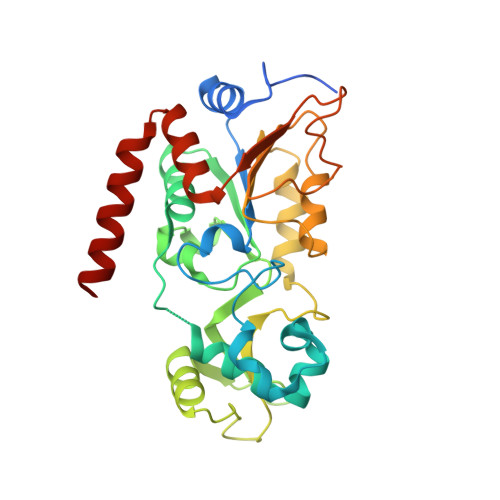
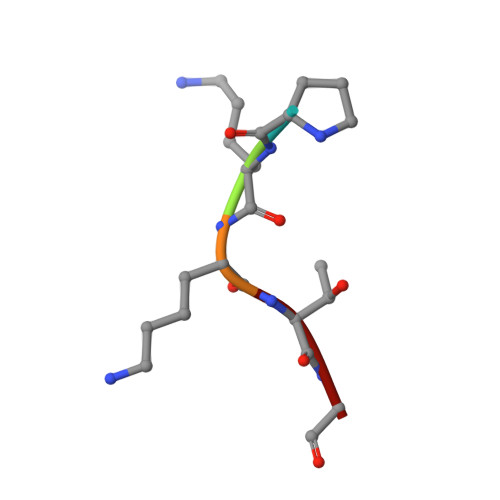
Efficient Demyristoylase Activity of SIRT2 Revealed by Kinetic and Structural Studies
Teng, Y.B., Jing, H., Aramsangtienchai, P., He, B., Khan, S., Hu, J., Lin, H., Hao, Q.(2015) Sci Rep 5: 8529-8529
- PubMed: 25704306
- DOI: https://doi.org/10.1038/srep08529
- Primary Citation of Related Structures:
4R8M - PubMed Abstract:
Sirtuins are a class of enzymes originally identified as nicotinamide adenine dinucleotide (NAD)-dependent protein lysine deacetylases. Among the seven mammalian sirtuins, SIRT1-7, only SIRT1-3 possess efficient deacetylase activity in vitro, whereas SIRT4-7 possess very weak in vitro deacetylase activity. Several sirtuins that exhibit weak deacetylase activity have recently been shown to possess more efficient activity for the removal other acyl lysine modifications, such as succinyl lysine and palmitoyl lysine. Here, we demonstrate that even the well-known deacetylase SIRT2 possesses efficient activity for the removal of long-chain fatty acyl groups. The catalytic efficiency (kcat/Km) for the removal of a myristoyl group is slightly higher than that for the removal of an acetyl group. The crystal structure of SIRT2 in complex with a thiomyristoyl peptide reveals that SIRT2 possesses a large hydrophobic pocket that can accommodate the myristoyl group. Comparison of the SIRT2 acyl pocket to those of SIRT1, SIRT3, and SIRT6 reveals that the acyl pockets of SIRT1-3 are highly similar, and to a lesser degree, similar to that of SIRT6. The efficient in vitro demyristoylase activity of SIRT2 suggests that this activity may be physiologically relevant and warrants future investigative studies.
Organizational Affiliation:
Department of Biomedical Sciences, Cornell University, Ithaca, NY 14853, USA.