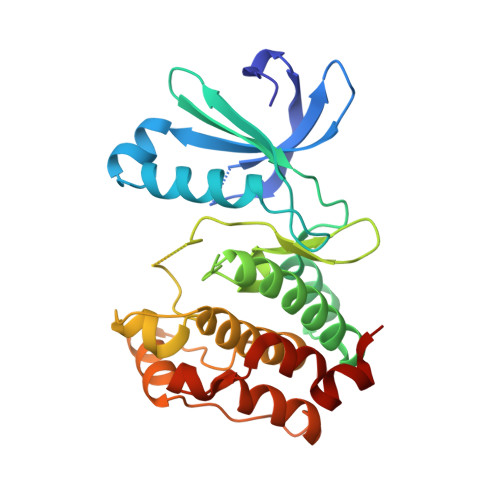
Property- and structure-guided discovery of a tetrahydroindazole series of interleukin-2 inducible T-cell kinase inhibitors.
Burch, J.D., Lau, K., Barker, J.J., Brookfield, F., Chen, Y., Chen, Y., Eigenbrot, C., Ellebrandt, C., Ismaili, M.H., Johnson, A., Kordt, D., MacKinnon, C.H., McEwan, P.A., Ortwine, D.F., Stein, D.B., Wang, X., Winkler, D., Yuen, P.W., Zhang, Y., Zarrin, A.A., Pei, Z.(2014) J Med Chem 57: 5714-5727
- PubMed: 24918870
- DOI: https://doi.org/10.1021/jm500550e
- Primary Citation of Related Structures:
4PQN, 4PRJ - PubMed Abstract:
Interleukin-2 inducible T-cell kinase (ITK), a member of the Tec family of tyrosine kinases, plays a major role in T-cell signaling downstream of the T-cell receptor (TCR), and considerable efforts have been directed toward discovery of ITK-selective inhibitors as potential treatments of inflammatory disorders such as asthma. Using a previously disclosed indazole series of inhibitors as a starting point, and using X-ray crystallography and solubility forecast index (SFI) as guides, we evolved a series of tetrahydroindazole inhibitors with improved potency, selectivity, and pharmaceutical properties. Highlights include identification of a selectivity pocket above the ligand plane, and identification of appropriate lipophilic substituents to occupy this space. This effort culminated in identification of a potent and selective ITK inhibitor (GNE-9822) with good ADME properties in preclinical species.
Organizational Affiliation:
Genentech Inc. , 1 DNA Way, South San Francisco, California 94080, United States.