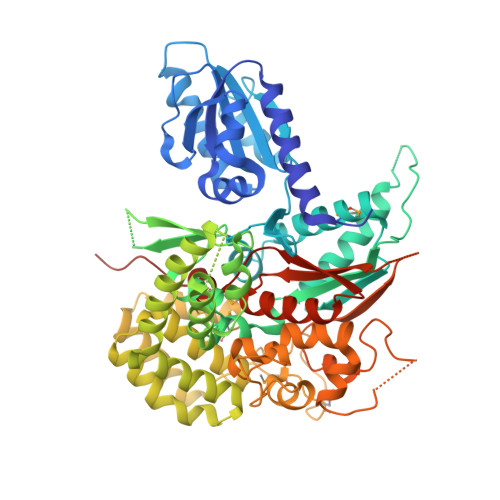
Crystal Structures of the Sec1/Munc18 (SM) Protein Vps33, Alone and Bound to the Homotypic Fusion and Vacuolar Protein Sorting (HOPS) Subunit Vps16*
Baker, R.W., Jeffrey, P.D., Hughson, F.M.(2013) PLoS One 8: e67409-e67409
- PubMed: 23840694
- DOI: https://doi.org/10.1371/journal.pone.0067409
- Primary Citation of Related Structures:
4JC8, 4KMO - PubMed Abstract:
Intracellular membrane fusion requires the regulated assembly of SNARE (soluble N-ethylmaleimide-sensitive factor (NSF) attachment protein receptor) proteins anchored in the apposed membranes. To exert the force required to drive fusion between lipid bilayers, juxtamembrane SNARE motifs zipper into four-helix bundles. Importantly, SNARE function is regulated by additional factors, none more extensively studied than the SM (Sec1/Munc18-like) proteins. SM proteins interact with both individual SNAREs and SNARE complexes, likely chaperoning SNARE complex formation and protecting assembly intermediates from premature disassembly by NSF. Four families of SM proteins have been identified, and representative members of two of these families (Sec1/Munc18 and Sly1) have been structurally characterized. We report here the 2.6 Å resolution crystal structure of an SM protein from the third family, Vps33. Although Vps33 shares with the first two families the same basic three-domain architecture, domain 1 is displaced by 15 Å, accompanied by a 40° rotation. A unique feature of the Vps33 family of SM proteins is that its members function as stable subunits within a multi-subunit tethering complex called HOPS (homotypic fusion and vacuolar protein sorting). Integration into the HOPS complex depends on the interaction between Vps33 and a second HOPS subunit, Vps16. The crystal structure of Vps33 bound to a C-terminal portion of Vps16, also at 2.6 Å resolution, reveals the structural basis for this interaction. Despite the extensive interface between the two HOPS subunits, the conformation of Vps33 is only subtly affected by binding to Vps16.
Organizational Affiliation:
Department of Molecular Biology, Princeton University, Princeton, New Jersey, United States of America.