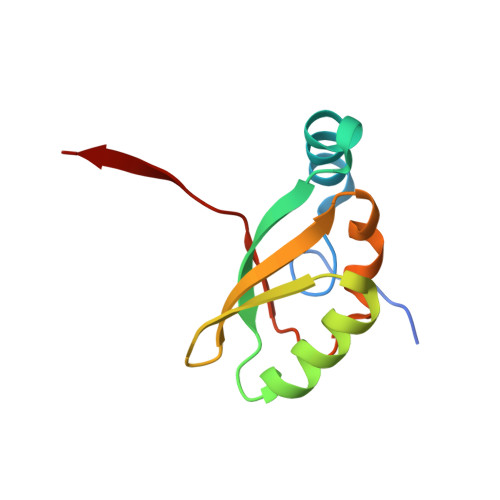
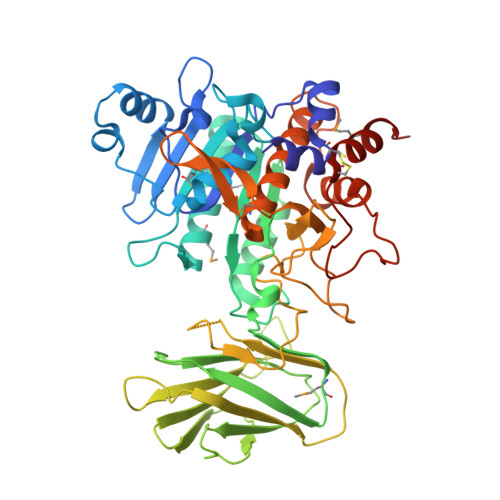
Structural and functional analysis of the CspB protease required for Clostridium spore germination.
Adams, C.M., Eckenroth, B.E., Putnam, E.E., Doublie, S., Shen, A.(2013) PLoS Pathog 9: e1003165-e1003165
- PubMed: 23408892
- DOI: https://doi.org/10.1371/journal.ppat.1003165
- Primary Citation of Related Structures:
4I0W - PubMed Abstract:
Spores are the major transmissive form of the nosocomial pathogen Clostridium difficile, a leading cause of healthcare-associated diarrhea worldwide. Successful transmission of C. difficile requires that its hardy, resistant spores germinate into vegetative cells in the gastrointestinal tract. A critical step during this process is the degradation of the spore cortex, a thick layer of peptidoglycan surrounding the spore core. In Clostridium sp., cortex degradation depends on the proteolytic activation of the cortex hydrolase, SleC. Previous studies have implicated Csps as being necessary for SleC cleavage during germination; however, their mechanism of action has remained poorly characterized. In this study, we demonstrate that CspB is a subtilisin-like serine protease whose activity is essential for efficient SleC cleavage and C. difficile spore germination. By solving the first crystal structure of a Csp family member, CspB, to 1.6 Å, we identify key structural domains within CspB. In contrast with all previously solved structures of prokaryotic subtilases, the CspB prodomain remains tightly bound to the wildtype subtilase domain and sterically occludes a catalytically competent active site. The structure, combined with biochemical and genetic analyses, reveals that Csp proteases contain a unique jellyroll domain insertion critical for stabilizing the protease in vitro and in C. difficile. Collectively, our study provides the first molecular insight into CspB activity and function. These studies may inform the development of inhibitors that can prevent clostridial spore germination and thus disease transmission.
Organizational Affiliation:
Graduate Program in Cell, Molecular and Biomedical Sciences, University of Vermont, Burlington, Vermont, United States of America.