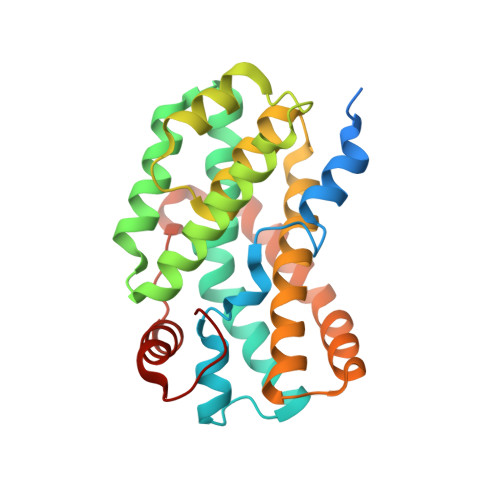
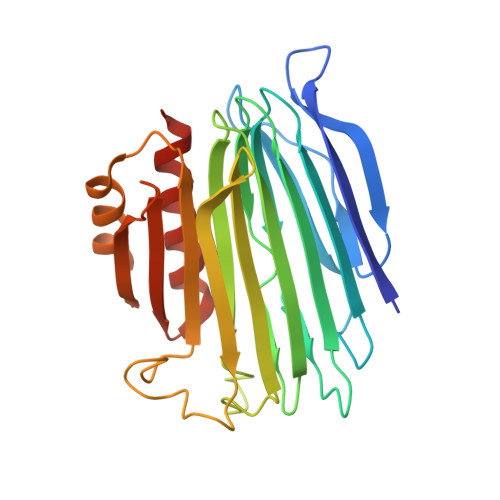
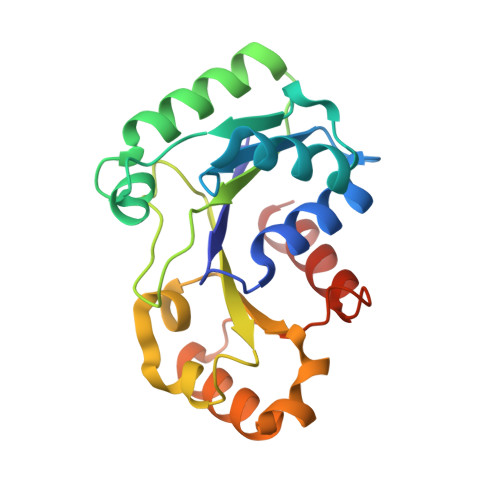
Structure of UreG/UreF/UreH complex reveals how urease accessory proteins facilitate maturation of Helicobacter pylori urease.
Fong, Y.H., Wong, H.C., Yuen, M.H., Lau, P.H., Chen, Y.W., Wong, K.B.(2013) PLoS Biol 11: e1001678-e1001678
- PubMed: 24115911
- DOI: https://doi.org/10.1371/journal.pbio.1001678
- Primary Citation of Related Structures:
4HI0 - PubMed Abstract:
Urease is a metalloenzyme essential for the survival of Helicobacter pylori in acidic gastric environment. Maturation of urease involves carbamylation of Lys219 and insertion of two nickel ions at its active site. This process requires GTP hydrolysis and the formation of a preactivation complex consisting of apo-urease and urease accessory proteins UreF, UreH, and UreG. UreF and UreH form a complex to recruit UreG, which is a SIMIBI class GTPase, to the preactivation complex. We report here the crystal structure of the UreG/UreF/UreH complex, which illustrates how UreF and UreH facilitate dimerization of UreG, and assembles its metal binding site by juxtaposing two invariant Cys66-Pro67-His68 metal binding motif at the interface to form the (UreG/UreF/UreH)2 complex. Interaction studies revealed that addition of nickel and GTP to the UreG/UreF/UreH complex releases a UreG dimer that binds a nickel ion at the dimeric interface. Substitution of Cys66 and His68 with alanine abolishes the formation of the nickel-charged UreG dimer. This nickel-charged UreG dimer can activate urease in vitro in the presence of the UreF/UreH complex. Static light scattering and atomic absorption spectroscopy measurements demonstrated that the nickel-charged UreG dimer, upon GTP hydrolysis, reverts to its monomeric form and releases nickel to urease. Based on our results, we propose a mechanism on how urease accessory proteins facilitate maturation of urease.
Organizational Affiliation:
School of Life Sciences, Center for Protein Science and Crystallography, The Chinese University of Hong Kong, Hong Kong, China.