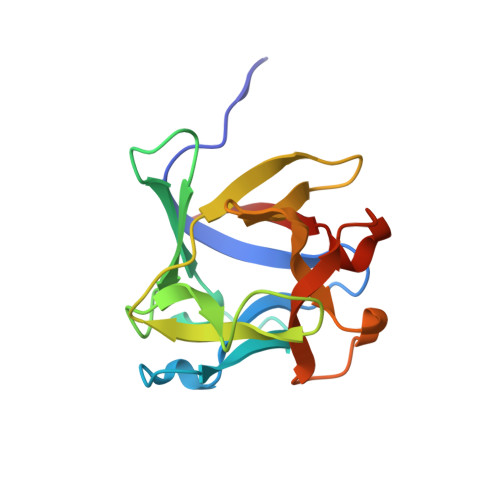
Structural analysis of the Rhizoctonia solani agglutinin reveals a domain-swapping dimeric assembly.
Skamnaki, V.T., Peumans, W.J., Kantsadi, A.L., Cubeta, M.A., Plas, K., Pakala, S., Zographos, S.E., Smagghe, G., Nierman, W.C., Van Damme, E.J., Leonidas, D.D.(2013) FEBS J 280: 1750-1763
- PubMed: 23402398
- DOI: https://doi.org/10.1111/febs.12190
- Primary Citation of Related Structures:
4G9M, 4G9N - PubMed Abstract:
Rhizoctonia solani agglutinin (RSA) is a 15.5-kDa lectin accumulated in the mycelium and sclerotia of the soil born plant pathogenic fungus R. solani. Although it is considered to serve as a storage protein and is implicated in fungal insecticidal activity, its physiological role remains unclear as a result of a lack of any structure/function relationship information. Glycan arrays showed that RSA displays high selectivity towards terminal nonreducing N-acetylgalactosamine residues. We determined the amino acid sequence of RSA and also determined the crystal structures of the free form and the RSA-N-acetylgalactosamine complex at 1.6 and 2.2 Å resolution, respectively. RSA is a homodimer comprised of two monomers adopting the β-trefoil fold. Each monomer accommodates two different carbohydrate-binding sites in an asymmetric way. Despite RSA topology similarities with R-type lectins, the two-monomer assembly involves an N-terminal swap, thus creating a dimer association novel to R-type lectins. Structural characterization of the two carbohydrate-binding sites offers insights on the structural determinants of the RSA carbohydrate specificity.
Organizational Affiliation:
Department of Biochemistry and Biotechnology, University of Thessaly, Larissa, Greece.