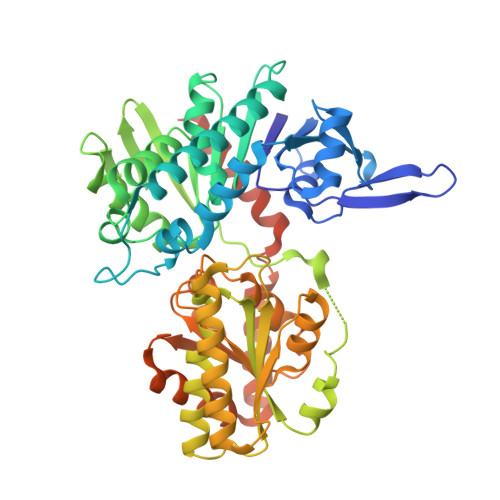
Mechanistic Insights into Validoxylamine A 7'-Phosphate Synthesis by VldE Using the Structure of the Entire Product Complex.
Cavalier, M.C., Yim, Y.S., Asamizu, S., Neau, D., Almabruk, K.H., Mahmud, T., Lee, Y.H.(2012) PLoS One 7: e44934-e44934
- PubMed: 23028689
- DOI: https://doi.org/10.1371/journal.pone.0044934
- Primary Citation of Related Structures:
4F96, 4F97, 4F9F - PubMed Abstract:
The pseudo-glycosyltransferase VldE catalyzes non-glycosidic C-N coupling between an unsaturated cyclitol and a saturated aminocyclitol with the conservation of the stereochemical configuration of the substrates to form validoxylamine A 7'-phosphate, the biosynthetic precursor of the antibiotic validamycin A. To study the molecular basis of its mechanism, the three-dimensional structures of VldE from Streptomyces hygroscopicus subsp. limoneus was determined in apo form, in complex with GDP, in complex with GDP and validoxylamine A 7'-phosphate, and in complex with GDP and trehalose. The structure of VldE with the catalytic site in both an "open" and "closed" conformation is also described. With these structures, the preferred binding of the guanine moiety by VldE, rather than the uracil moiety as seen in OtsA could be explained. The elucidation of the VldE structure in complex with the entirety of its products provides insight into the internal return mechanism by which catalysis occurs with a net retention of the stereochemical configuration of the donated cyclitol.
Organizational Affiliation:
Department of Biological Sciences, Louisiana State University, Baton Rouge, Louisiana, United States of America.