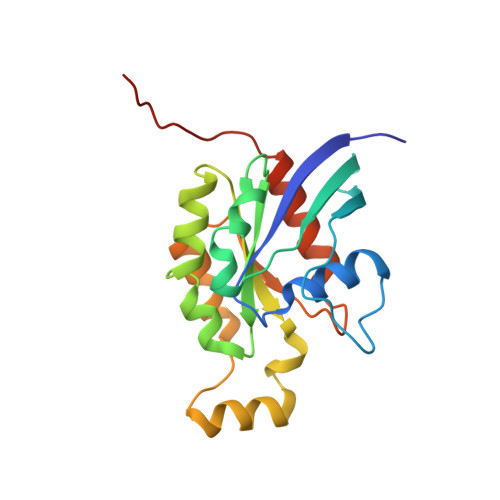
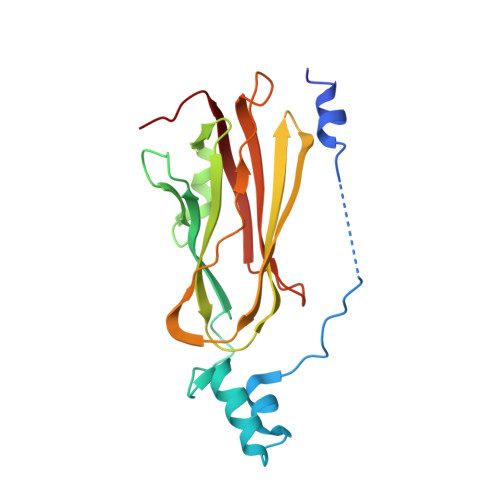
Quantitative Analysis of Prenylated RhoA Interaction with Its Chaperone, RhoGDI.
Tnimov, Z., Guo, Z., Gambin, Y., Nguyen, U.T., Wu, Y.W., Abankwa, D., Stigter, A., Collins, B.M., Waldmann, H., Goody, R.S., Alexandrov, K.(2012) J Biol Chem 287: 26549-26562
- PubMed: 22628549
- DOI: https://doi.org/10.1074/jbc.M112.371294
- Primary Citation of Related Structures:
4F38 - PubMed Abstract:
Small GTPases of the Rho family regulate cytoskeleton remodeling, cell polarity, and transcription, as well as the cell cycle, in eukaryotic cells. Membrane delivery and recycling of the Rho GTPases is mediated by Rho GDP dissociation inhibitor (RhoGDI), which forms a stable complex with prenylated Rho GTPases. We analyzed the interaction of RhoGDI with the active and inactive forms of prenylated and unprenylated RhoA. We demonstrate that RhoGDI binds the prenylated form of RhoA·GDP with unexpectedly high affinity (K(d) = 5 pm). The very long half-life of the complex is reduced 25-fold on RhoA activation, with a concomitant reduction in affinity (K(d) = 3 nm). The 2.8-Å structure of the RhoA·guanosine 5'-[β,γ-imido] triphosphate (GMPPNP)·RhoGDI complex demonstrated that complex formation forces the activated RhoA into a GDP-bound conformation in the absence of nucleotide hydrolysis. We demonstrate that membrane extraction of Rho GTPase by RhoGDI is a thermodynamically favored passive process that operates through a series of progressively tighter intermediates, much like the one that is mediated by RabGDI.
Organizational Affiliation:
Department of Molecular Cell Biology, Institute for Molecular Bioscience, The University of Queensland, 306 Carmody Road, St. Lucia, Queensland 4072, Australia.