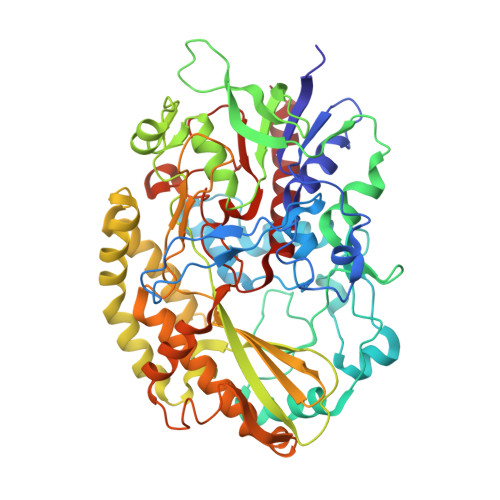
Structural analysis of fungus-derived FAD glucose dehydrogenase
Yoshida, H., Sakai, G., Mori, K., Kojima, K., Kamitori, S., Sode, K.(2015) Sci Rep 5: 13498-13498
- PubMed: 26311535
- DOI: https://doi.org/10.1038/srep13498
- Primary Citation of Related Structures:
4YNT, 4YNU - PubMed Abstract:
We report the first three-dimensional structure of fungus-derived glucose dehydrogenase using flavin adenine dinucleotide (FAD) as the cofactor. This is currently the most advanced and popular enzyme used in glucose sensor strips manufactured for glycemic control by diabetic patients. We prepared recombinant nonglycosylated FAD-dependent glucose dehydrogenase (FADGDH) derived from Aspergillus flavus (AfGDH) and obtained the X-ray structures of the binary complex of enzyme and reduced FAD at a resolution of 1.78 Å and the ternary complex with reduced FAD and D-glucono-1,5-lactone (LGC) at a resolution of 1.57 Å. The overall structure is similar to that of fungal glucose oxidases (GOxs) reported till date. The ternary complex with reduced FAD and LGC revealed the residues recognizing the substrate. His505 and His548 were subjected for site-directed mutagenesis studies, and these two residues were revealed to form the catalytic pair, as those conserved in GOxs. The absence of residues that recognize the sixth hydroxyl group of the glucose of AfGDH, and the presence of significant cavity around the active site may account for this enzyme activity toward xylose. The structural information will contribute to the further engineering of FADGDH for use in more reliable and economical biosensing technology for diabetes management.
Organizational Affiliation:
Life Science Research Center and Faculty of Medicine, 1750-1, Ikenobe, Miki-cho, Kita-gun, Kagawa University, Kagawa 761-0793, Japan.