Cooperative folding of intrinsically disordered domains drives assembly of a strong elongated protein.
Gruszka, D.T., Whelan, F., Farrance, O.E., Fung, H.K., Paci, E., Jeffries, C.M., Svergun, D.I., Baldock, C., Baumann, C.G., Brockwell, D.J., Potts, J.R., Clarke, J.(2015) Nat Commun 6: 7271-7271
- PubMed: 26027519
- DOI: https://doi.org/10.1038/ncomms8271
- Primary Citation of Related Structures:
4WVE - PubMed Abstract:
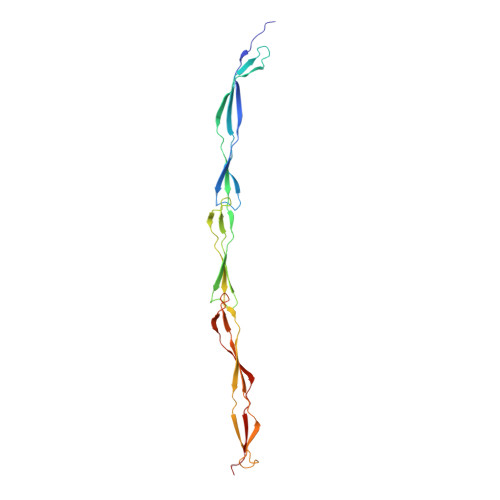
Bacteria exploit surface proteins to adhere to other bacteria, surfaces and host cells. Such proteins need to project away from the bacterial surface and resist significant mechanical forces. SasG is a protein that forms extended fibrils on the surface of Staphylococcus aureus and promotes host adherence and biofilm formation. Here we show that although monomeric and lacking covalent cross-links, SasG maintains a highly extended conformation in solution. This extension is mediated through obligate folding cooperativity of the intrinsically disordered E domains that couple non-adjacent G5 domains thermodynamically, forming interfaces that are more stable than the domains themselves. Thus, counterintuitively, the elongation of the protein appears to be dependent on the inherent instability of its domains. The remarkable mechanical strength of SasG arises from tandemly arrayed 'clamp' motifs within the folded domains. Our findings reveal an elegant minimal solution for the assembly of monomeric mechano-resistant tethers of variable length.
Organizational Affiliation:
Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge CB2 1EW, UK.