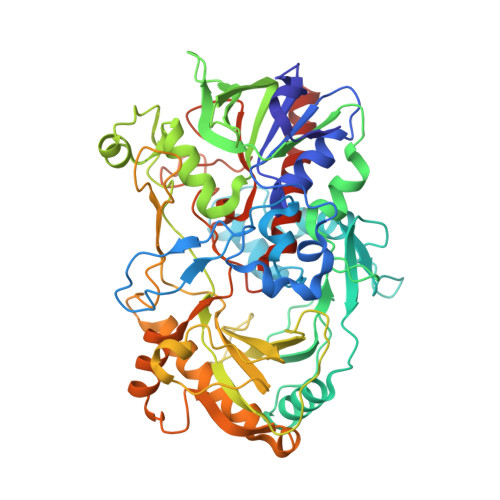
Structure of choline oxidase in complex with the reaction product glycine betaine.
Salvi, F., Wang, Y.F., Weber, I.T., Gadda, G.(2014) Acta Crystallogr D Biol Crystallogr 70: 405-413
- PubMed: 24531474
- DOI: https://doi.org/10.1107/S1399004713029283
- Primary Citation of Related Structures:
4MJW - PubMed Abstract:
Choline oxidase from Arthrobacter globiformis, which is involved in the biosynthesis of glycine betaine from choline, has been extensively characterized in its mechanistic and structural properties. Despite the knowledge gained on the enzyme, the details of substrate access to the active site are not fully understood. The `loop-and-lid' mechanism described for the glucose-methanol-choline enzyme superfamily has not been confirmed for choline oxidase. Instead, a hydrophobic cluster on the solvent-accessible surface of the enzyme has been proposed by molecular dynamics to control substrate access to the active site. Here, the crystal structure of the enzyme was solved in complex with glycine betaine at pH 6.0 at 1.95 Å resolution, allowing a structural description of the ligand-enzyme interactions in the active site. This structure is the first of choline oxidase in complex with a physiologically relevant ligand. The protein structures with and without ligand are virtually identical, with the exception of a loop at the dimer interface, which assumes two distinct conformations. The different conformations of loop 250-255 define different accessibilities of the proposed active-site entrance delimited by the hydrophobic cluster on the other subunit of the dimer, suggesting a role in regulating substrate access to the active site.
Organizational Affiliation:
Department of Chemistry, Georgia State University, PO Box 3965, Atlanta, GA 30302-3965, USA.