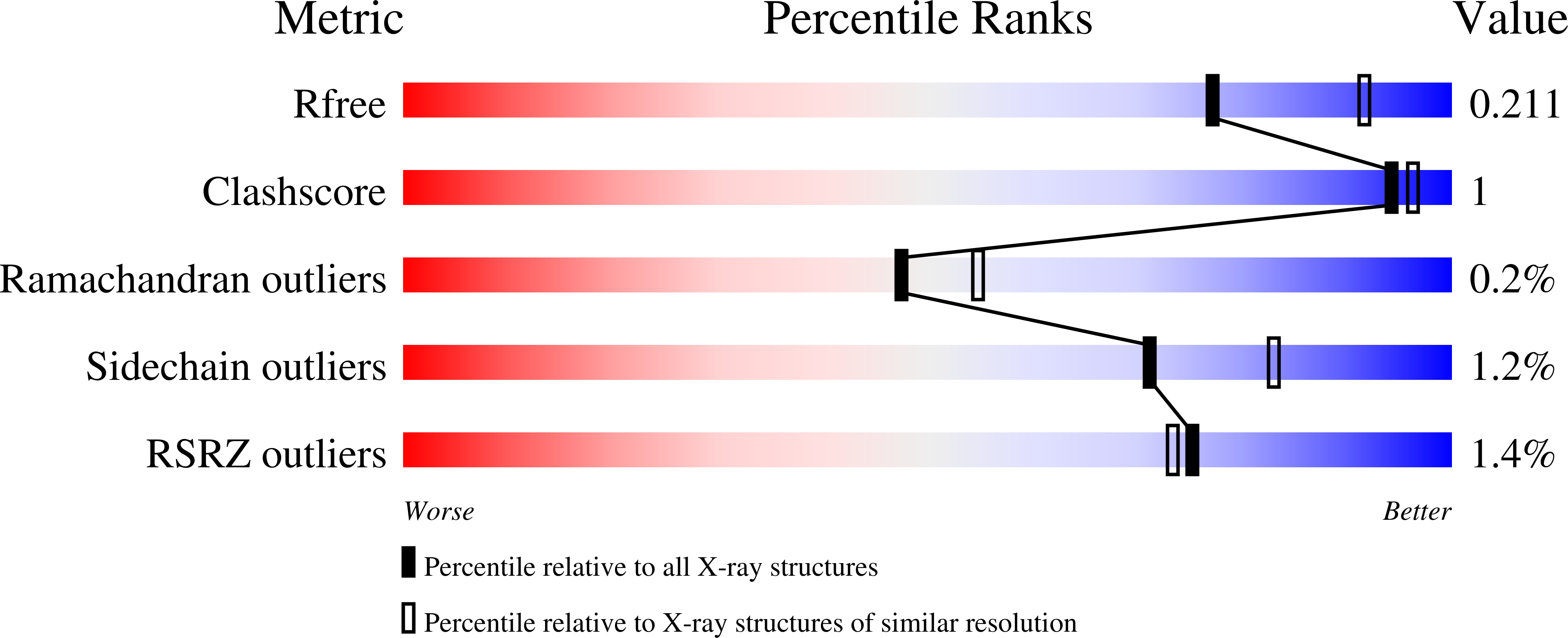
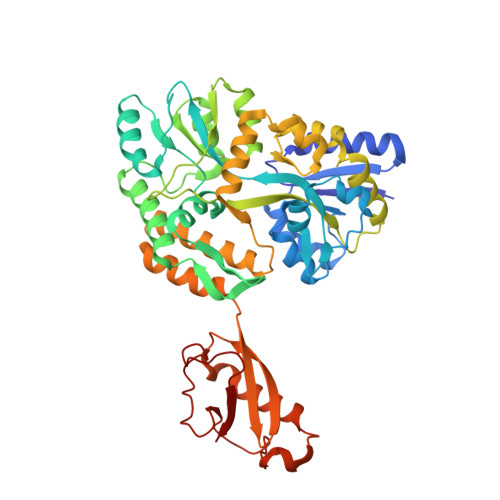
Structures of EccB1 and EccD1 from the core complex of the mycobacterial ESX-1 type VII secretion system.
Wagner, J.M., Chan, S., Evans, T.J., Kahng, S., Kim, J., Arbing, M.A., Eisenberg, D., Korotkov, K.V.(2016) BMC Struct Biol 16: 5-5
- PubMed: 26922638
- DOI: https://doi.org/10.1186/s12900-016-0056-6
- Primary Citation of Related Structures:
4KK7, 4KV2, 4KV3, 5CYU - PubMed Abstract:
The ESX-1 type VII secretion system is an important determinant of virulence in pathogenic mycobacteria, including Mycobacterium tuberculosis. This complicated molecular machine secretes folded proteins through the mycobacterial cell envelope to subvert the host immune response. Despite its important role in disease very little is known about the molecular architecture of the ESX-1 secretion system. This study characterizes the structures of the soluble domains of two conserved core ESX-1 components - EccB1 and EccD1. The periplasmic domain of EccB1 consists of 4 repeat domains and a central domain, which together form a quasi 2-fold symmetrical structure. The repeat domains of EccB1 are structurally similar to a known peptidoglycan binding protein suggesting a role in anchoring the ESX-1 system within the periplasmic space. The cytoplasmic domain of EccD1has a ubiquitin-like fold and forms a dimer with a negatively charged groove. These structures represent a major step towards resolving the molecular architecture of the entire ESX-1 assembly and may contribute to ESX-1 targeted tuberculosis intervention strategies.
Organizational Affiliation:
Department of Molecular & Cellular Biochemistry and Center for Structural Biology, University of Kentucky, 741 South Limestone, Lexington, KY, 40536, USA. jonw7782@gmail.com.