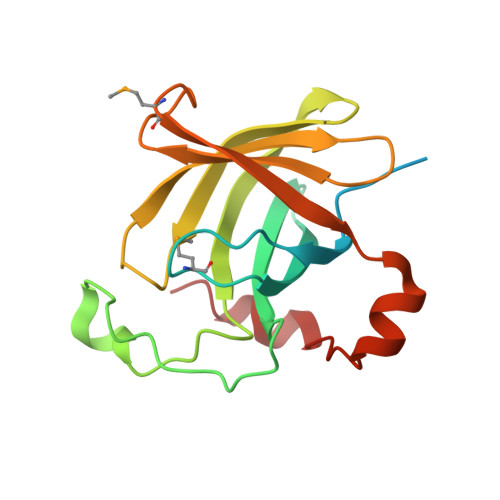
Protein HP1028 from the human pathogen Helicobacter pylori belongs to the lipocalin family.
Barison, N., Cendron, L., Loconte, V., Proctor, E.A., Dokholyan, N.V., Zanotti, G.(2013) Acta Crystallogr D Biol Crystallogr 69: 1387-1394
- PubMed: 23897462
- DOI: https://doi.org/10.1107/S0907444913008160
- Primary Citation of Related Structures:
4INN - PubMed Abstract:
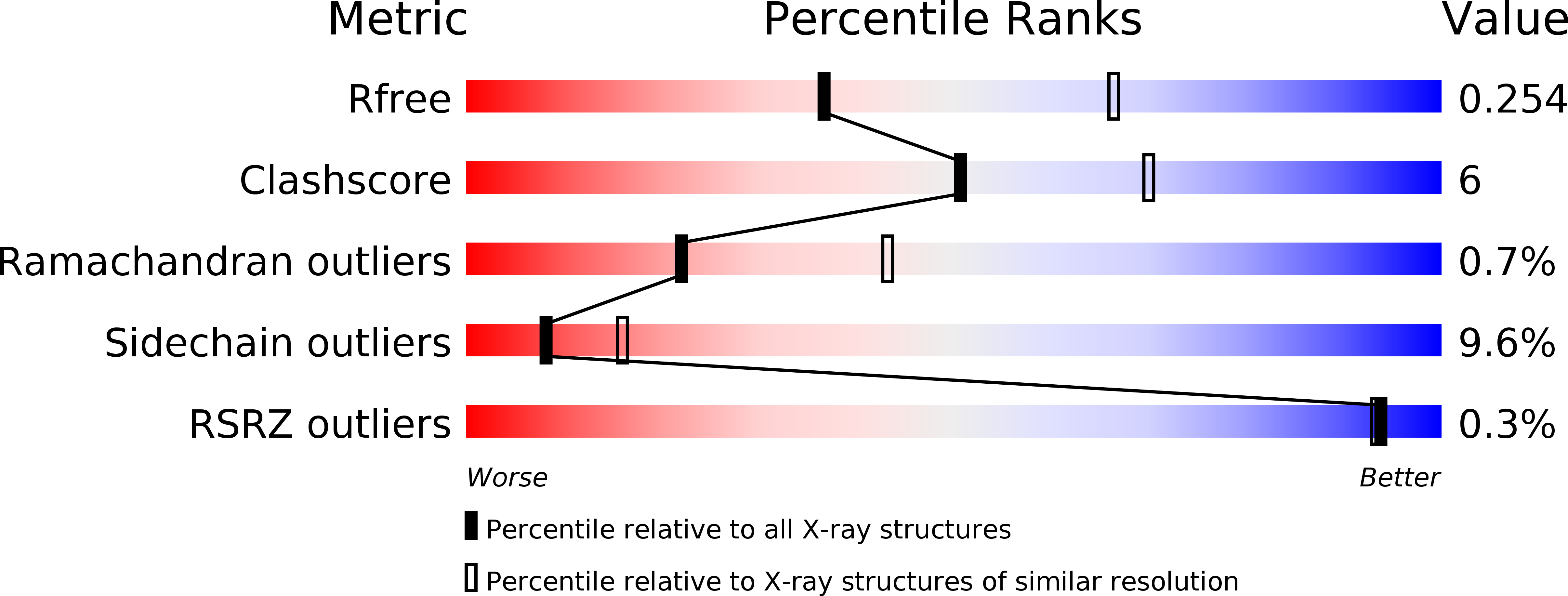
Helicobacter pylori is a bacterial pathogen that causes severe diseases, including gastritis, ulcers and gastric cancer. Although this bacterium has been extensively studied, the physiological functions of a large number of the proteins encoded by its genome are unknown. HP1028 is a protein that is relevant to colonization and to the survival of the bacterium in the stomach, but its function is not clearly understood. Bioinformatics studies suggest that HP1028 is a monomeric protein that is secreted in the H. pylori periplasm. The crystal structure of HP1028 has been determined at 2.6 Å resolution using the SAD method. The three-dimensional structure of the protein reveals that it belongs to the lipocalin family, a group of proteins that bind and transport (often hydrophobic) small molecules. The structure of HP1028, together with the possible localization of the mature protein in the bacterial periplasm and the position of the hp1028 gene in the bacterial genome, point to a role in H. pylori chemotaxis.
Organizational Affiliation:
Department of Biomedical Sciences, University of Padua, Viale G. Colombo 3, 35131 Padova, Italy.