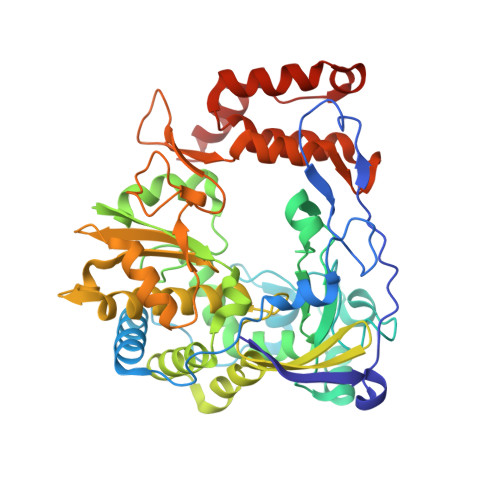
Crystal structure of enterovirus 71 RNA-dependent RNA polymerase complexed with its protein primer VPg: implication for a trans mechanism of VPg uridylylation
Chen, C., Wang, Y.X., Shan, C., Sun, Y., Xu, P., Zhou, H.G., Yang, C., Shi, P.Y., Rao, Z.H., Zhang, B., Lou, Z.Y.(2013) J Virol 87: 5755-5768
- PubMed: 23487447
- DOI: https://doi.org/10.1128/JVI.02733-12
- Primary Citation of Related Structures:
4IKA - PubMed Abstract:
Picornavirus RNA replication is initiated by VPg uridylylation, during which the hydroxyl group of the third tyrosine residue of the virally encoded protein VPg is covalently linked to two UMP molecules by RNA-dependent RNA polymerase (RdRp; also known as 3D(pol)). We previously identified site 311, located at the base of the palm domain of the enterovirus 71 (EV71) RdRp, to be the site for EV71 VPg binding and uridylylation. Here we report the crystal structure of EV71 3D(pol) complexed with VPg. VPg was anchored at the bottom of the palm domain of the 3D(pol) molecule and exhibited an extended V-shape conformation. The corresponding interface on 3D(pol) was mainly formed by residues within site 311 and other residues in the palm and finger domains. Mutations of the amino acids of 3D(pol) involved in the VPg interaction (3DL319A, 3DD320A, and 3DY335A) significantly disrupted VPg binding to 3D(pol), resulting in defective VPg uridylylation. In contrast, these mutations did not affect the RNA elongation activity of 3D(pol). In the context of viral genomic RNA, mutations that abolished VPg uridylylation activity were lethal for EV71 replication. Further in vitro analysis showed that the uridylylation activity was restored by mixing VPg-binding-defective and catalysis-defective mutants, indicating a trans mechanism for EV71 VPg uridylylation. Our results, together with previous results of other studies, demonstrate that different picornaviruses use distinct binding sites for VPg uridylylation.
Organizational Affiliation:
Structural Biology Laboratory and MOE Laboratory of Protein Science, School of Medicine and Life Science, Tsinghua University, Beijing, China.