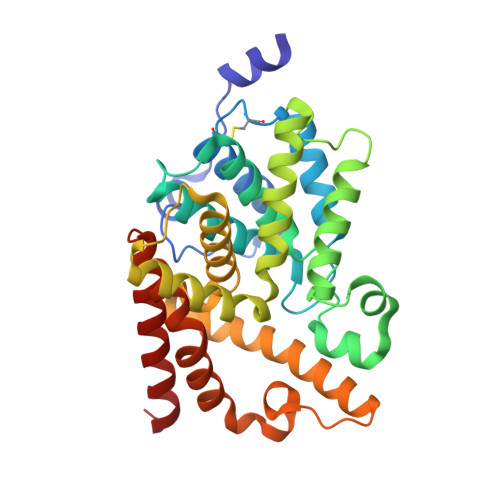
Discovery of selective biaryl ethers as PDE10A inhibitors: Improvement in potency and mitigation of Pgp-mediated efflux.
Rzasa, R.M., Hu, E., Rumfelt, S., Chen, N., Andrews, K.L., Chmait, S., Falsey, J.R., Zhong, W., Jones, A.D., Porter, A., Louie, S.W., Zhao, X., Treanor, J.J., Allen, J.R.(2012) Bioorg Med Chem Lett 22: 7371-7375
- PubMed: 23149228
- DOI: https://doi.org/10.1016/j.bmcl.2012.10.078
- Primary Citation of Related Structures:
4HEU, 4HF4 - PubMed Abstract:
We report the discovery of a novel series of biaryl ethers as potent and selective PDE10A inhibitors. Structure-activity studies improved the potency and decreased Pgp-mediated efflux found in the initial compound 4. X-ray crystallographic studies revealed two novel binding modes to the catalytic site of the PDE10A enzyme.
Organizational Affiliation:
Department of Medicinal Chemistry, Amgen Inc., One Amgen Center Drive, Thousand Oaks, CA 91320-1799, USA. rrzasa@amgen.com