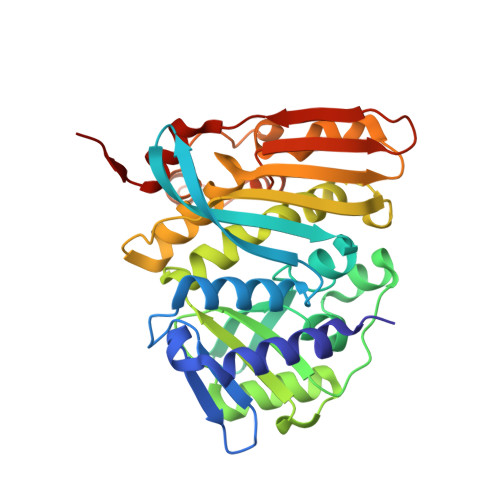
Structure and catalytic mechanism of N(5),N(10)-methenyl-tetrahydromethanopterin cyclohydrolase.
Upadhyay, V., Demmer, U., Warkentin, E., Moll, J., Shima, S., Ermler, U.(2012) Biochemistry 51: 8435-8443
- PubMed: 23013430
- DOI: https://doi.org/10.1021/bi300777k
- Primary Citation of Related Structures:
4GVQ, 4GVR, 4GVS - PubMed Abstract:
Methenyltetrahydromethanopterin (methenyl-H(4)MPT(+)) cyclohydrolase (Mch) catalyzes the interconversion of methenyl-H(4)MPT(+) and formyl-H(4)MPT in the one-carbon energy metabolism of methanogenic, methanotrophic, and sulfate-reducing archaea and of methylotrophic bacteria. To understand the catalytic mechanism of this reaction, we kinetically characterized site-specific variants of Mch from Archaeoglobus fulgidus (aMch) and determined the X-ray structures of the substrate-free aMch(E186Q), the aMch:H(4)MPT complex, and the aMch(E186Q):formyl-H(4)MPT complex. (Formyl-)H(4)MPT is embedded inside a largely preformed, interdomain pocket of the homotrimeric enzyme with the pterin and benzyl rings being oriented nearly perpendicular to each other. The active site is primarily built up by the segment 93:95, Arg183 and Glu186 that either interact with the catalytic water attacking methenyl-H(4)MPT(+) or with the formyl oxygen of formyl-H(4)MPT. The catalytic function of the strictly conserved Arg183 and Glu186 was substantiated by the low enzymatic activities of the E186A, E186D, E186N, E186Q, R183A, R183Q, R183E, R183K, and R183E-E186Q variants. Glu186 most likely acts as a general base. Arg183 decisively influences the pK(a) value of Glu186 and the proposed catalytic water mainly by its positive charge. In addition, Glu186 appears to be also responsible for product specificity by donating a proton to the directly neighbored N(10) tertiary amine of H(4)MPT. Thus, N(10) becomes a better leaving group than N(5) which implies the generation of N(5)-formyl-H(4)MPT. For comparison, methenyltetrahydrofolate (H(4)F) cyclohydrolase produces N(10)-formyl-H(4)F in an analogous reaction. An enzymatic mechanism of Mch is postulated and compared with that of other cyclohydrolases.
Organizational Affiliation:
Max-Planck-Institut für Biophysik, Max-von-Laue-Straße 3, D-60438 Frankfurt am Main, Germany.