Modular Architecture and Unique Teichoic Acid Recognition Features of Choline-Binding Protein L (Cbpl) Contributing to Pneumococcal Pathogenesis.
Gutierrez-Fernandez, J., Saleh, M., Alcorlo, M., Gomez-Mejia, A., Pantoja-Uceda, D., Trevino, M.A., Vos, F., Abdullah, M.R., Galan-Bartual, S., Seinen, J., Sanchez-Murcia, P.A., Gago, F., Bruix, M., Hammerschmidt, S., Hermoso, J.A.(2016) Sci Rep 6: 38094
- PubMed: 27917891
- DOI: https://doi.org/10.1038/srep38094
- Primary Citation of Related Structures:
4CNL - PubMed Abstract:
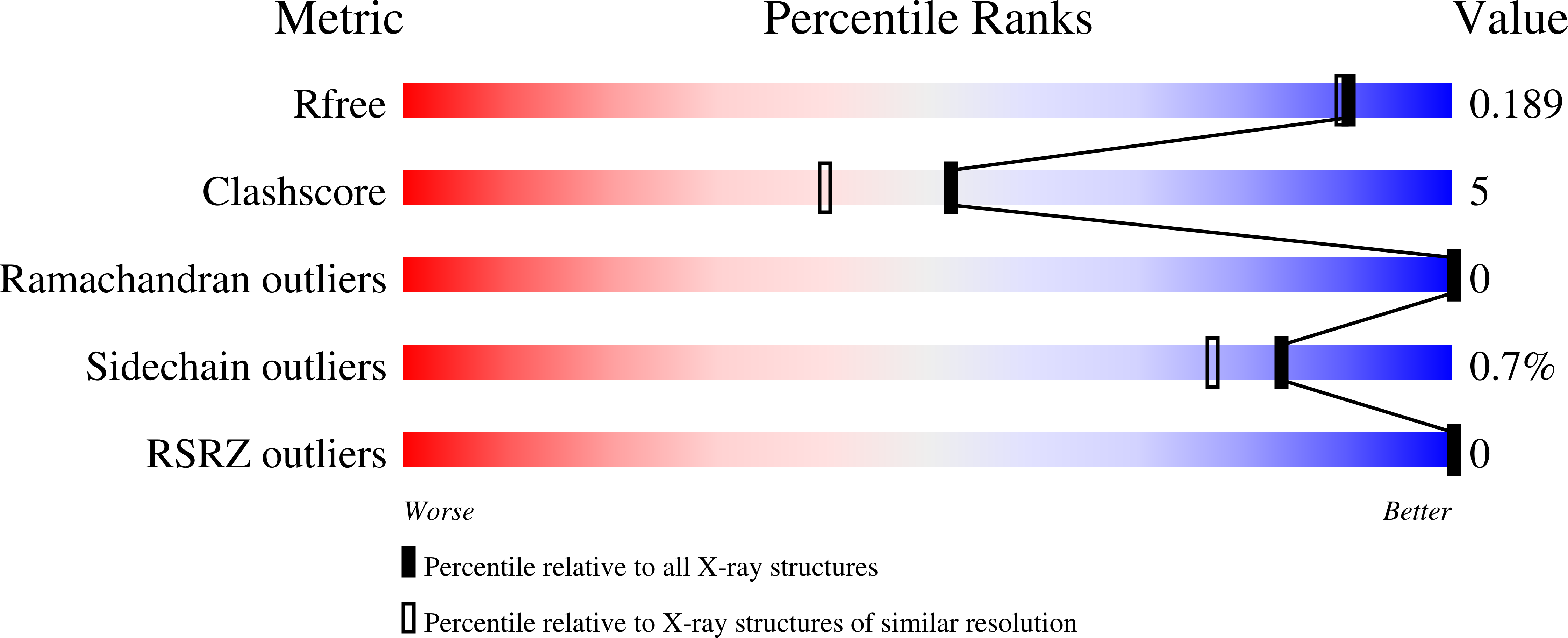
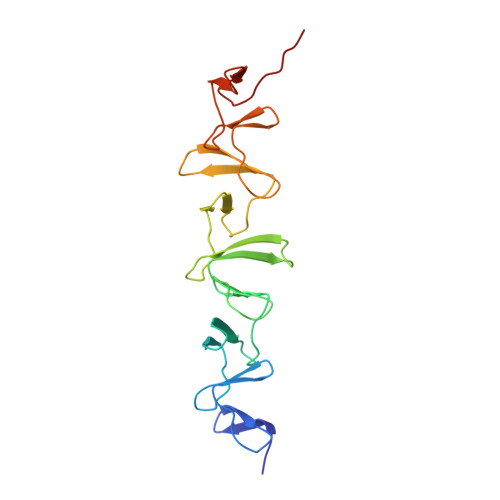
The human pathogen Streptococcus pneumoniae is decorated with a special class of surface-proteins known as choline-binding proteins (CBPs) attached to phosphorylcholine (PCho) moieties from cell-wall teichoic acids. By a combination of X-ray crystallography, NMR, molecular dynamics techniques and in vivo virulence and phagocytosis studies, we provide structural information of choline-binding protein L (CbpL) and demonstrate its impact on pneumococcal pathogenesis and immune evasion. CbpL is a very elongated three-module protein composed of (i) an Excalibur Ca 2+ -binding domain -reported in this work for the very first time-, (ii) an unprecedented anchorage module showing alternate disposition of canonical and non-canonical choline-binding sites that allows vine-like binding of fully-PCho-substituted teichoic acids (with two choline moieties per unit), and (iii) a Ltp_Lipoprotein domain. Our structural and infection assays indicate an important role of the whole multimodular protein allowing both to locate CbpL at specific places on the cell wall and to interact with host components in order to facilitate pneumococcal lung infection and transmigration from nasopharynx to the lungs and blood. CbpL implication in both resistance against killing by phagocytes and pneumococcal pathogenesis further postulate this surface-protein as relevant among the pathogenic arsenal of the pneumococcus.
Organizational Affiliation:
Department of Crystallography and Structural Biology, "Rocasolano" Institute of Physical-Chemistry, CSIC, Serrano 119, E-28006-Madrid, Spain.