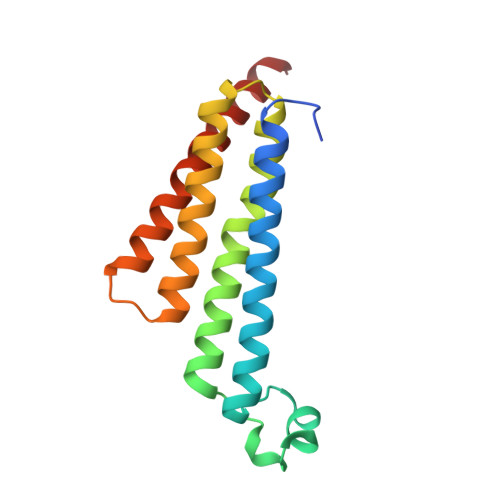
Crystal Structure of Microsomal Prostaglandin E2 Synthase Provides Insight Into Diversity in the Mapeg Superfamily.
Sjogren, T., Nord, J., Ek, M., Johansson, P., Liu, G., Geschwindner, S.(2013) Proc Natl Acad Sci U S A 110: 3806
- PubMed: 23431194
- DOI: https://doi.org/10.1073/pnas.1218504110
- Primary Citation of Related Structures:
4AL0, 4AL1 - PubMed Abstract:
Prostaglandin E2 (PGE2) is a key mediator in inflammatory response. The main source of inducible PGE2, microsomal PGE2 synthase-1 (mPGES-1), has emerged as an interesting drug target for treatment of pain. To support inhibitor design, we have determined the crystal structure of human mPGES-1 to 1.2 Å resolution. The structure reveals three well-defined active site cavities within the membrane-spanning region in each monomer interface of the trimeric structure. An important determinant of the active site cavity is a small cytosolic domain inserted between transmembrane helices I and II. This extra domain is not observed in other structures of proteins within the MAPEG (Membrane-Associated Proteins involved in Eicosanoid and Glutathione metabolism) superfamily but is likely to be present also in microsomal GST-1 based on sequence similarity. An unexpected feature of the structure is a 16-Å-deep cone-shaped cavity extending from the cytosolic side into the membrane-spanning region. We suggest a potential role for this cavity in substrate access. Based on the structure of the active site, we propose a catalytic mechanism in which serine 127 plays a key role. We have also determined the structure of mPGES-1 in complex with a glutathione-based analog, providing insight into mPGES-1 flexibility and potential for structure-based drug design.
Organizational Affiliation:
Discovery Sciences, AstraZeneca R&D Mölndal, S-43183 Mölndal, Sweden. tove.sjogren@astrazeneca.com