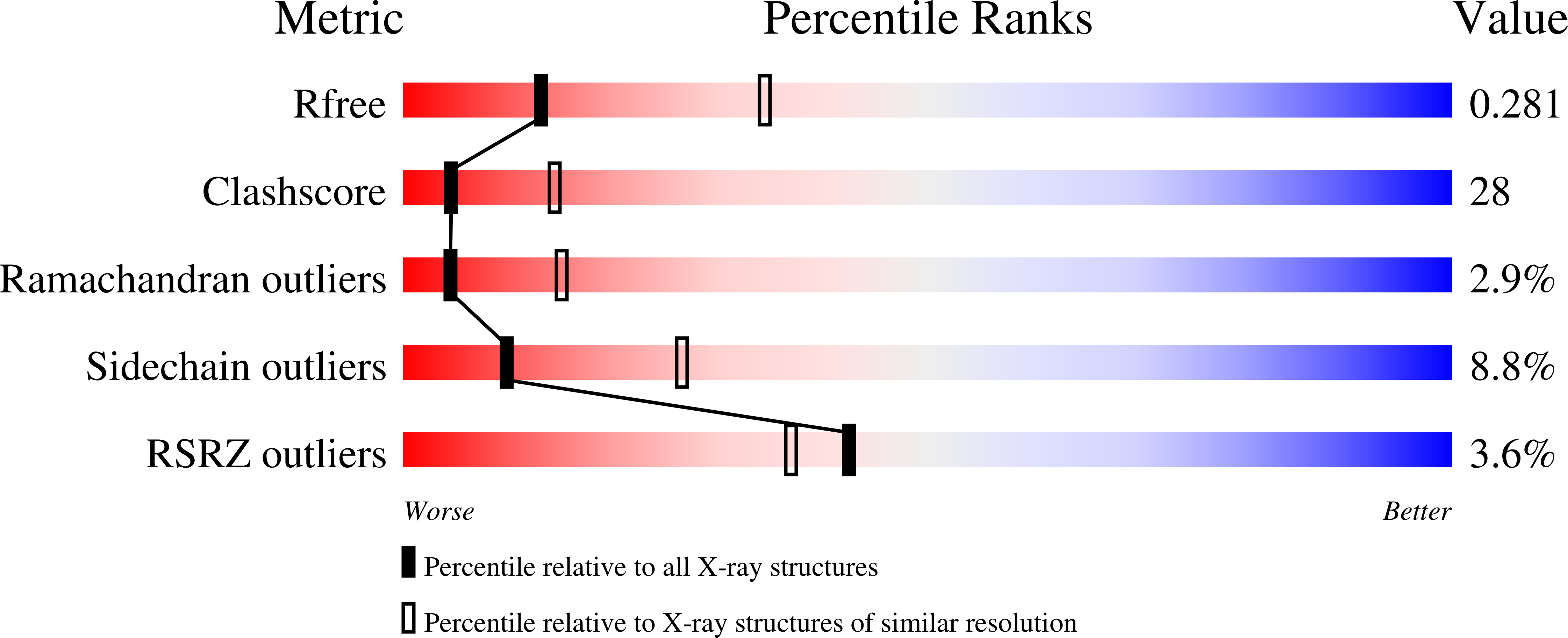
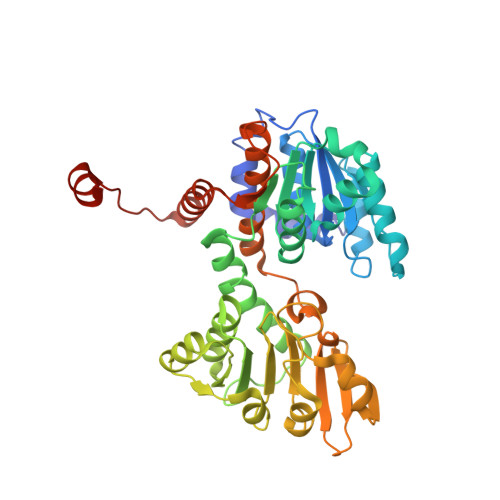
Crystal structures of S-adenosylhomocysteine hydrolase from the thermophilic bacterium Thermotoga maritima.
Zheng, Y., Chen, C.C., Ko, T.P., Xiao, X., Yang, Y., Huang, C.H., Qian, G., Shao, W., Guo, R.T.(2015) J Struct Biol 190: 135-142
- PubMed: 25791616
- DOI: https://doi.org/10.1016/j.jsb.2015.03.002
- Primary Citation of Related Structures:
3X2E, 3X2F - PubMed Abstract:
S-adenosylhomocysteine (SAH) hydrolase catalyzes the reversible hydrolysis of SAH into adenosine and homocysteine by using NAD(+) as a cofactor. The enzyme from Thermotoga maritima (tmSAHH) has great potentials in industrial applications because of its hyperthermophilic properties. Here, two crystal structures of tmSAHH in complex with NAD(+) show both open and closed conformations despite the absence of bound substrate. Each subunit of the tetrameric enzyme is composed of three domains, namely the catalytic domain, the NAD(+)-binding domain and the C-terminal domain. The NAD(+) binding mode is clearly observed and a substrate analogue can also be modeled into the active site, where two cysteine residues in mesophilic enzymes are replaced by serine and threonine in tmSAHH. Notably, the C-terminal domain of tmSAHH lacks the second loop region of mesophilic SAHH, which is important in NAD(+) binding, and thus exposes the bound cofactor to the solvent. The difference explains the higher NAD(+) requirement of tmSAHH because of the reduced affinity. Furthermore, the feature of missing loop is consistently observed in thermophilic bacterial and archaeal SAHHs, and may be related to their thermostability.
Organizational Affiliation:
Industrial Enzymes National Engineering Laboratory, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin 300308, China.