Biophysical characterization of mutants of Bacillus subtilis lipase evolved for thermostability: Factors contributing to increased activity retention.
Augustyniak, W., Brzezinska, A.A., Pijning, T., Wienk, H., Boelens, R., Dijkstra, B.W., Reetz, M.T.(2012) Protein Sci 21: 487-497
- PubMed: 22267088
- DOI: https://doi.org/10.1002/pro.2031
- Primary Citation of Related Structures:
3QZU - PubMed Abstract:
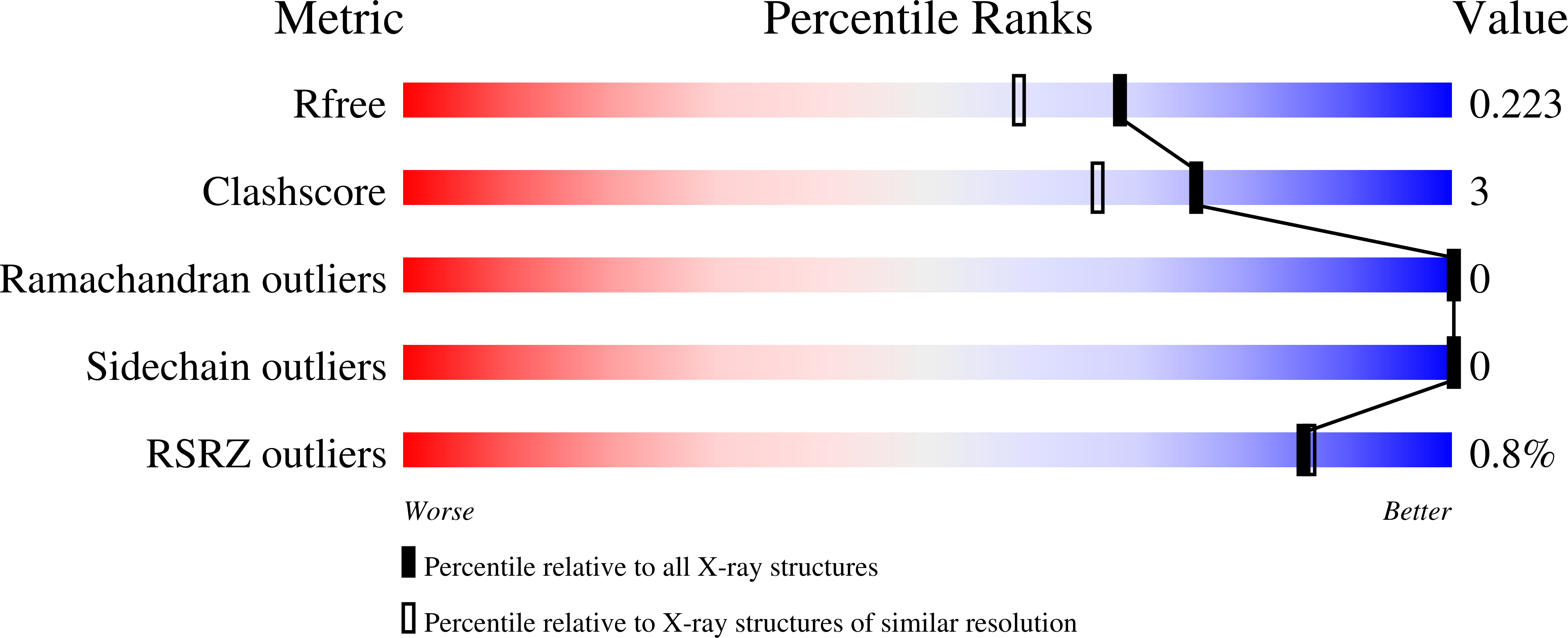
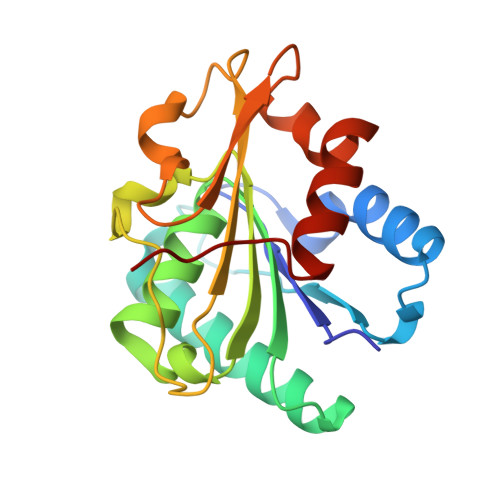
Previously, Lipase A from Bacillus subtilis was subjected to in vitro directed evolution using iterative saturation mutagenesis, with randomization sites chosen on the basis of the highest B-factors available from the crystal structure of the wild-type (WT) enzyme. This provided mutants that, unlike WT enzyme, retained a large part of their activity after heating above 65 °C and cooling down. Here, we subjected the three best mutants along with the WT enzyme to biophysical and biochemical characterization. Combining thermal inactivation profiles, circular dichroism, X-ray structure analyses and NMR experiments revealed that mutations of surface amino acid residues counteract the tendency of Lipase A to undergo precipitation under thermal stress. Reduced precipitation of the unfolding intermediates rather than increased conformational stability of the evolved mutants seems to be responsible for the activity retention.
Organizational Affiliation:
Max-Planck-Institut fur Kohlenforschung, Kaiser-Wilhelm-Platz 1, 45470 Mulheim an der Ruhr, Germany.