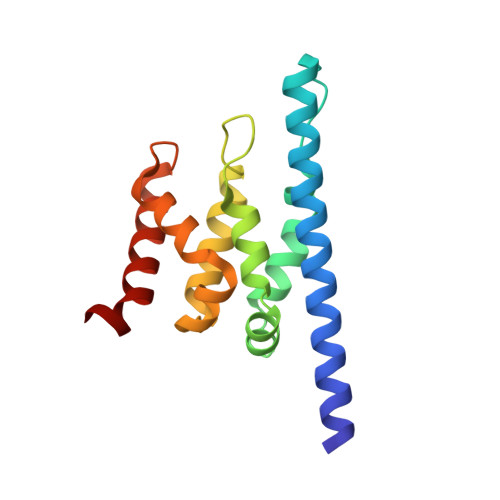
Structure and biological importance of the spn1-spt6 interaction, and its regulatory role in nucleosome binding.
McDonald, S.M., Close, D., Xin, H., Formosa, T., Hill, C.P.(2010) Mol Cell 40: 725-735
- PubMed: 21094070
- DOI: https://doi.org/10.1016/j.molcel.2010.11.014
- Primary Citation of Related Structures:
3O8Z, 3OAK - PubMed Abstract:
Eukaryotic transcription and mRNA processing depend upon the coordinated interactions of many proteins, including Spn1 and Spt6, which are conserved across eukaryotes, are essential for viability, and associate with each other in some of their biologically important contexts. Here we report crystal structures of the Spn1 core alone and in complex with the binding determinant of Spt6. Mutating interface residues greatly diminishes binding in vitro and causes strong phenotypes in vivo, including a defect in maintaining repressive chromatin. Overexpression of Spn1 partially suppresses the defects caused by an spt6 mutation affecting the Spn1 interface, indicating that the Spn1-Spt6 interaction is important for managing chromatin. Spt6 binds nucleosomes directly in vitro, and this interaction is blocked by Spn1, providing further mechanistic insight into the function of the interaction. These data thereby reveal the structural and biochemical bases of molecular interactions that function in the maintenance of chromatin structure.
Organizational Affiliation:
Department of Biochemistry, University of Utah School of Medicine, Salt Lake City, UT 84112-5650, USA.