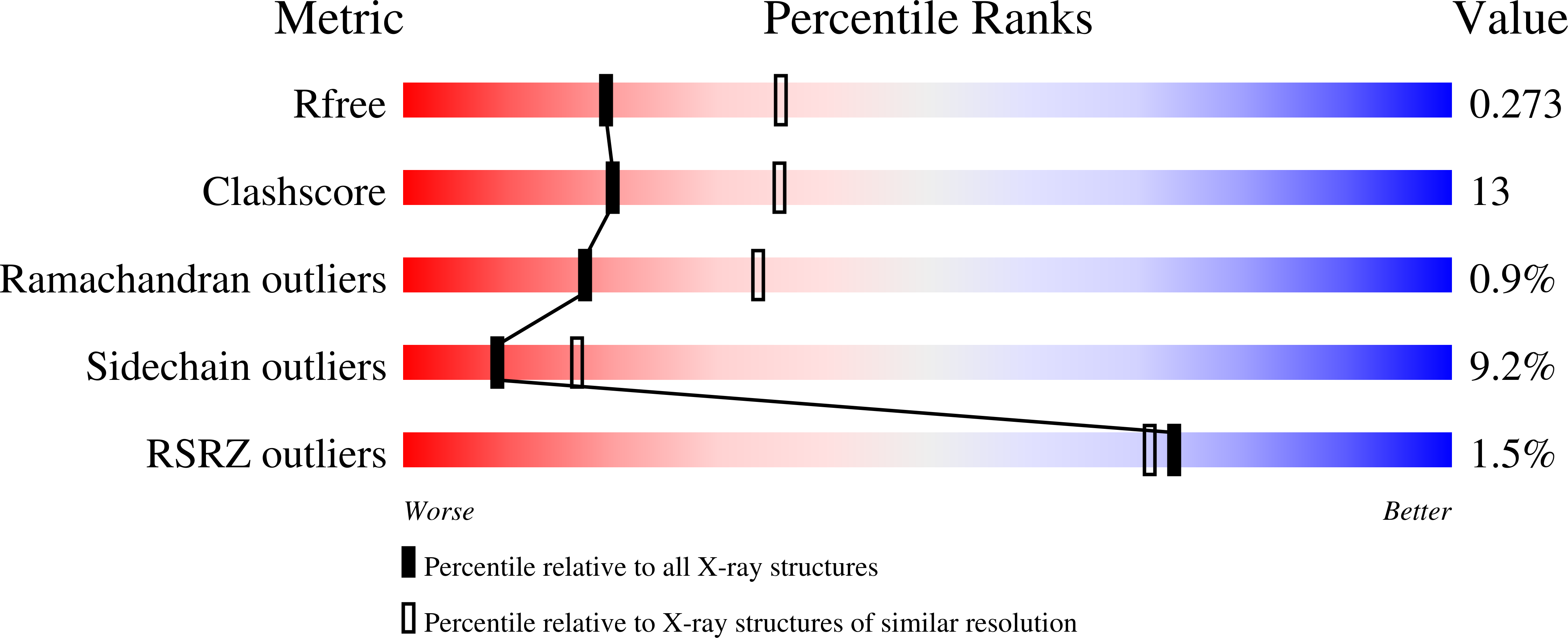
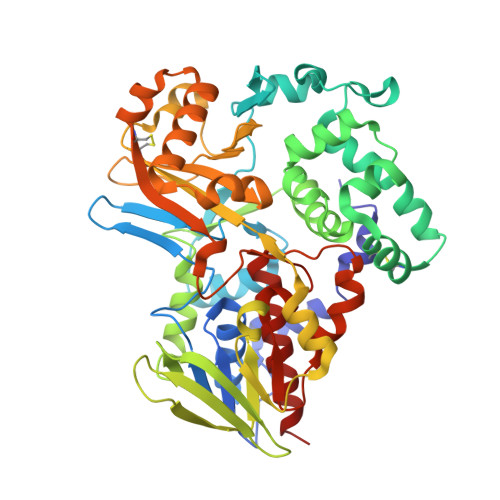
Structure of native L-amino acid oxidase from Vipera ammodytes ammodytes: stabilization of the quaternary structure by divalent ions and structural changes in the dynamic active site
Gergiova, D., Murakami, M.T., Perbandt, M., Arni, R.K., Betzel, C.To be published.