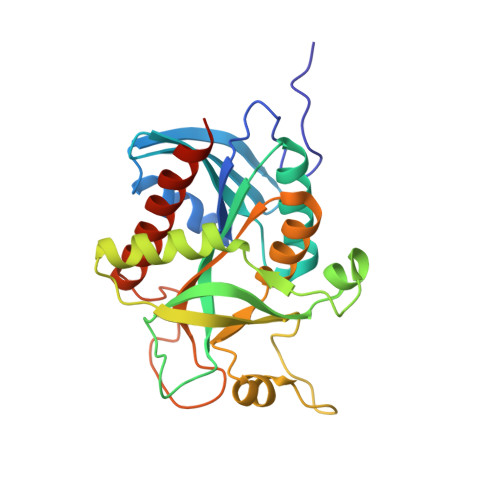
Conservation of structure and activity in Plasmodium purine nucleoside phosphorylases
Chaikuad, A., Brady, R.L.(2009) BMC Struct Biol 9: 42-42
- PubMed: 19575810
- DOI: https://doi.org/10.1186/1472-6807-9-42
- Primary Citation of Related Structures:
3EMV, 3ENZ - PubMed Abstract:
Purine nucleoside phosphorylase (PNP) is central to purine salvage mechanisms in Plasmodium parasites, the causative agents of malaria. Most human malaria results from infection either by Plasmodium falciparum (Pf), the deadliest form of the parasite, or by the widespread Plasmodium vivax (Pv). Whereas the PNP enzyme from Pf has previously been studied in detail, despite the prevalence of Pv little is known about many of the key metabolic enzymes from this parasite, including PvPNP.
Organizational Affiliation:
Department of Biochemistry, University of Bristol, Bristol, BS8 1TD, UK. Apirat.Chaikuad@sgc.ox.ac.uk