Crystal Structure of Full Length Circadian Clock Protein KaiC with Correct Geometry at Phosphorylation Sites
Pattanayek, R., Williams, D.R., Pattanayek, S., Xu, Y., Mori, T., Johnson, C.H., Stewart, P.L., Egli, M.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Circadian clock protein kinase kaiC | 519 | Synechococcus elongatus PCC 7942 = FACHB-805 | Mutation(s): 0 Gene Names: kaiC EC: 2.7.11.1 | 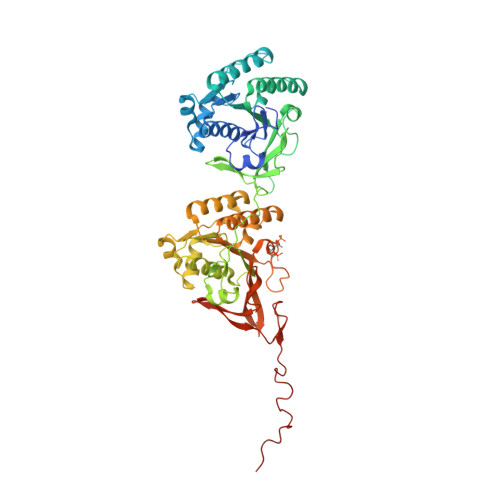 | |
UniProt | |||||
Find proteins for Q79PF4 (Synechococcus elongatus (strain ATCC 33912 / PCC 7942 / FACHB-805)) Explore Q79PF4 Go to UniProtKB: Q79PF4 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q79PF4 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ATP Query on ATP | H [auth A] I [auth A] K [auth B] L [auth B] N [auth C] | ADENOSINE-5'-TRIPHOSPHATE C10 H16 N5 O13 P3 ZKHQWZAMYRWXGA-KQYNXXCUSA-N |  | ||
| MG Query on MG | G [auth A] J [auth B] M [auth C] P [auth D] S [auth E] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| SEP Query on SEP | A, B, C, D, E A, B, C, D, E, F | L-PEPTIDE LINKING | C3 H8 N O6 P |  | SER |
| TPO Query on TPO | A, B, C, D, E A, B, C, D, E, F | L-PEPTIDE LINKING | C4 H10 N O6 P |  | THR |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 132.873 | α = 90 |
| b = 135.576 | β = 90 |
| c = 204.951 | γ = 90 |
| Software Name | Purpose |
|---|---|
| RAVE | model building |
| GLRF | phasing |
| CNS | refinement |
| DENZO | data reduction |
| SCALEPACK | data scaling |
| RAVE | phasing |