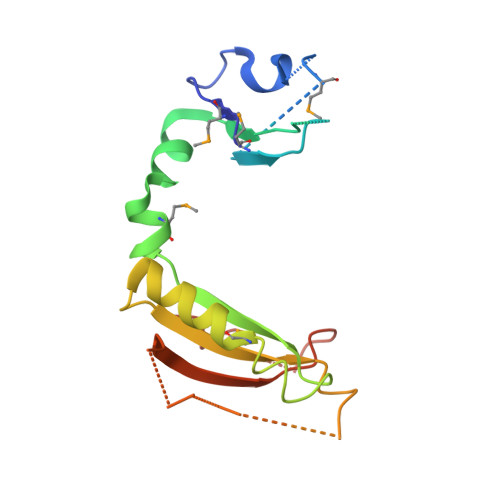
Structure of the ROC domain from the Parkinson's disease-associated leucine-rich repeat kinase 2 reveals a dimeric GTPase
Deng, J., Lewis, P.A., Greggio, E., Sluch, E., Beilina, A., Cookson, M.R.(2008) Proc Natl Acad Sci U S A 105: 1499-1504
- PubMed: 18230735
- DOI: https://doi.org/10.1073/pnas.0709098105
- Primary Citation of Related Structures:
2ZEJ, 3D6T - PubMed Abstract:
Mutations in leucine-rich repeat kinase 2 (LRRK2) are the most common cause of Parkinson's disease (PD). LRRK2 contains a Ras of complex proteins (ROC) domain that may act as a GTPase to regulate its protein kinase activity. The structure of ROC and the mechanism(s) by which it regulates kinase activity are not known. Here, we report the crystal structure of the LRRK2 ROC domain in complex with GDP-Mg(2+) at 2.0-A resolution. The structure displays a dimeric fold generated by extensive domain-swapping, resulting in a pair of active sites constructed with essential functional groups contributed from both monomers. Two PD-associated pathogenic residues, R1441 and I1371, are located at the interface of two monomers and provide exquisite interactions to stabilize the ROC dimer. The structure demonstrates that loss of stabilizing forces in the ROC dimer is likely related to decreased GTPase activity resulting from mutations at these sites. Our data suggest that the ROC domain may regulate LRRK2 kinase activity as a dimer, possibly via the C-terminal of ROC (COR) domain as a molecular hinge. The structure of the LRRK2 ROC domain also represents a signature from a previously undescribed class of GTPases from complex proteins and results may provide a unique molecular target for therapeutics in PD.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Oklahoma State University, Stillwater, OK 74078, USA. junpeng.deng@okstate.edu