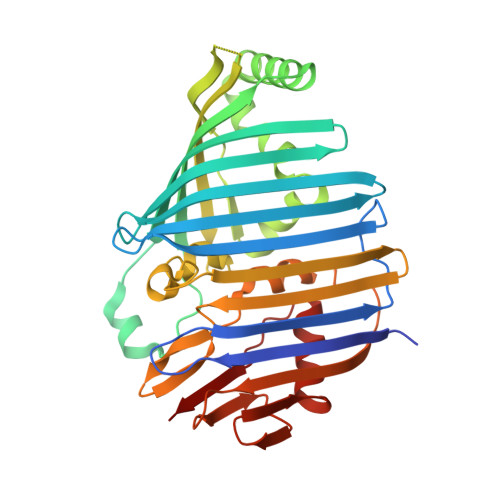
Reinterpretation of the electron density at the site of the eighth bacteriochlorophyll in the FMO protein from Pelodictyon phaeum.
Tronrud, D.E., Allen, J.P.(2012) Photosynth Res 112: 71-74
- PubMed: 22457093
- DOI: https://doi.org/10.1007/s11120-012-9735-8
- Primary Citation of Related Structures:
3VDI - PubMed Abstract:
The Fenna-Matthews-Olson antenna protein from the green bacterium Pelodictyon phaeum mediates the energy transfer from a peripheral antenna complex to the membrane-bound reaction center. The three-dimensional structure of this protein has been previously modeled using X-ray diffraction to a resolution limit of 2.0 Å, with R (work) and R (free) values of 16.6 and 19.9%, respectively (Larson et al., Photosynth Res 107:139-150, 2011). This model shows the protein as consisting of β-sheets surrounding several bacteriochlorophyll cofactors. While most of the model clearly matches the electron density maps, in this paper we re-examine the electron density for a specific feature, namely the eighth bacteriochlorophyll a cofactor. This electron density is now interpreted as arising primarily from the end of an otherwise disordered polyethylene glycol molecule. Additional electron density is present but the density is weak and cannot be unambiguously assigned. The new model has R (work) and R (free) values of 16.2 and 19.0%, respectively.
Organizational Affiliation:
Department of Biochemistry and Biophysics, Oregon State University, Corvallis, OR 97331, USA. tronrudd@science.oregonstate.edu