Evaluating the therapeutic potential of a non-natural nucleotide that inhibits human ribonucleotide reductase.
Ahmad, M.F., Wan, Q., Jha, S., Motea, E., Berdis, A., Dealwis, C.(2012) Mol Cancer Ther 11: 2077-2086
- PubMed: 22933704
- DOI: https://doi.org/10.1158/1535-7163.MCT-12-0199
- Primary Citation of Related Structures:
3RSR - PubMed Abstract:
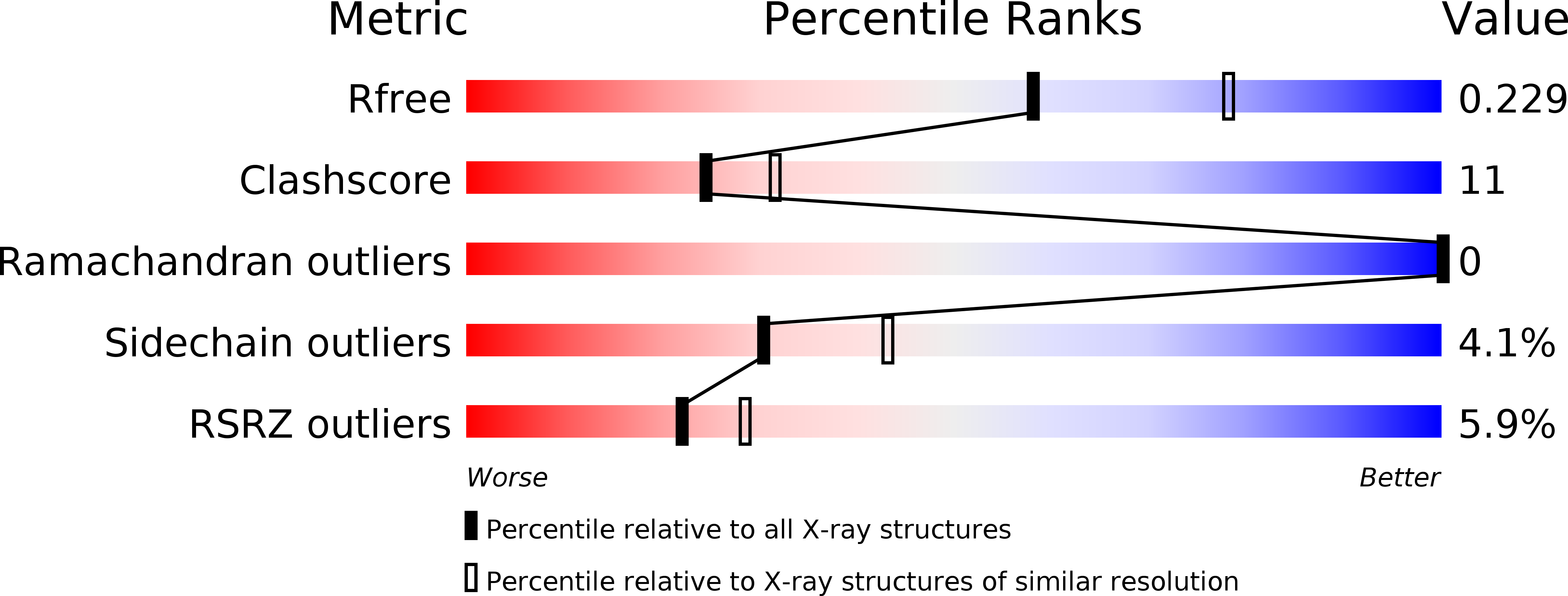
Human ribonucleotide reductase (hRR) is the key enzyme involved in de novo dNTP synthesis and thus represents an important therapeutic target against hyperproliferative diseases, most notably cancer. The purpose of this study was to evaluate the ability of non-natural indolyl-2'-deoxynucleoside triphosphates to inhibit the activity of hRR. The structural similarities of these analogues with dATP predicted that they would inhibit hRR activity by binding to its allosteric sites. In silico analysis and in vitro characterization identified one particular analogue designated as 5-nitro-indolyl-2'-deoxyribose triphosphate (5-NITP) that inhibits hRR. 5-NITP binding to hRR was determined by isothermal titration calorimetry. X-ray crystal structure of 5-NITP bound to RR1 was determined. Cell-based studies showed the anti-cancer effects of the corresponding non-natural nucleoside against leukemia cells. 5-NITP binds to hRR with micromolar affinity. Binding does not induce hexamerization of hRR1 like dATP, the native allosteric inhibitor of hRR that binds with high affinity to the A-site. The X-ray crystal structure of Saccharomyces cerevisiae RR1-5-NITP (ScRR1-5-NITP) complex determined to 2.3 Å resolution shows that 5-NITP does not bind to the A-site but rather at the S-site. Regardless, 5-nitro-indolyl-2'-deoxynucleoside (5-NIdR) produces cytostatic and cytotoxic effects against human leukemia cells by altering cell-cycle progression. Our studies provide useful insights toward developing new inhibitors with improved potency and efficacy against hRR.
Organizational Affiliation:
Corresponding Author: Chris Dealwis, Case Western Reserve University, 10900 Euclid Avenue, Wood Building, W303, Cleveland, OH 44106, USA.