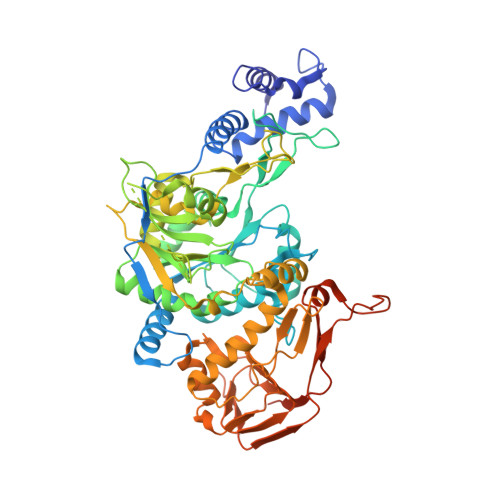
Structural basis for the broad substrate range of the UDP-sugar pyrophosphorylase from Leishmania major.
Dickmanns, A., Damerow, S., Neumann, P., Schulz, E.C., Lamerz, A.C., Routier, F.H., Ficner, R.(2011) J Mol Biol 405: 461-478
- PubMed: 21073876
- DOI: https://doi.org/10.1016/j.jmb.2010.10.057
- Primary Citation of Related Structures:
3OGZ, 3OH0, 3OH1, 3OH2, 3OH3, 3OH4 - PubMed Abstract:
Nucleotide sugars and the enzymes that are responsible for their synthesis are indispensable for the production of complex carbohydrates and, thus, for elaboration of a protective cellular coat for many organisms such as the protozoan parasite Leishmania. These activated sugars are synthesized de novo or derived from salvaged monosaccharides. In addition to UDP-glucose (UDP-Glc) pyrophosphorylase, which catalyzes the formation of UDP-Glc from substrates UTP and glucose-1-phosphate, Leishmania major and plants express a UDP-sugar pyrophosphorylase (USP) that exhibits broad substrate specificity in vitro. The enzyme, likely involved in monosaccharide salvage, preferentially generates UDP-Glc and UDP-galactose, but it may also activate other hexose- or pentose-1-phosphates such as galacturonic acid-1-phosphate or arabinose-1-phosphate. In order to gain insight into structural features governing the differences in substrate specificity, we determined the crystal structure of the L. major USP in the APO-, UTP-, and UDP-sugar-bound conformations. The overall tripartite structure of USP exhibits a significant structural homology to other nucleotidyldiphosphate-glucose pyrophosphorylases. The obtained USP structures reveal the structural rearrangements occurring during the stepwise binding process of the substrates. Moreover, the different product complexes explain the broad substrate specificity of USP, which is enabled by structural changes in the sugar binding region of the active site.
Organizational Affiliation:
Institut für Mikrobiologie und Genetik & GZMB, Georg-August-Universität Göttingen, Justus-von-Liebig-Weg 11, D-37077 Göttingen, Germany.