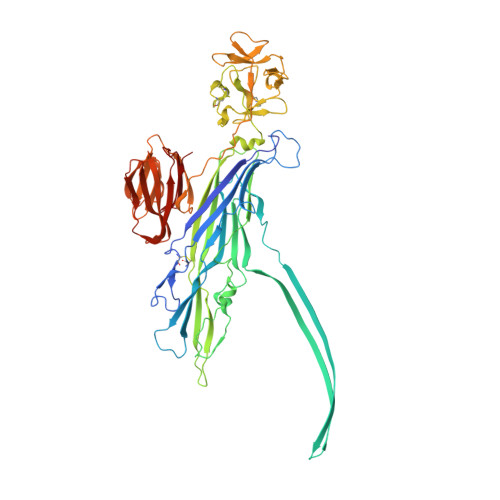
Crystal structure of the Vibrio cholerae cytolysin heptamer reveals common features among disparate pore-forming toxins.
De, S., Olson, R.(2011) Proc Natl Acad Sci U S A 108: 7385-7390
- PubMed: 21502531
- DOI: https://doi.org/10.1073/pnas.1017442108
- Primary Citation of Related Structures:
3O44 - PubMed Abstract:
Pore-forming toxins (PFTs) are potent cytolytic agents secreted by pathogenic bacteria that protect microbes against the cell-mediated immune system (by targeting phagocytic cells), disrupt epithelial barriers, and liberate materials necessary to sustain growth and colonization. Produced by gram-positive and gram-negative bacteria alike, PFTs are released as water-soluble monomeric or dimeric species, bind specifically to target membranes, and assemble transmembrane channels leading to cell damage and/or lysis. Structural and biophysical analyses of individual steps in the assembly pathway are essential to fully understanding the dynamic process of channel formation. To work toward this goal, we solved by X-ray diffraction the 2.9-Å structure of the 450-kDa heptameric Vibrio cholerae cytolysin (VCC) toxin purified and crystallized in the presence of detergent. This structure, together with our previously determined 2.3-Å structure of the VCC water-soluble monomer, reveals in detail the architectural changes that occur within the channel region and accessory lectin domains during pore formation including substantial rearrangements of hydrogen-bonding networks in the pore-forming amphipathic loops. Interestingly, a ring of tryptophan residues forms the narrowest constriction in the transmembrane channel reminiscent of the phenylalanine clamp identified in anthrax protective antigen [Krantz BA, et al. (2005) Science 309:777-781]. Our work provides an example of a β-barrel PFT (β-PFT) for which soluble and assembled structures are available at high-resolution, providing a template for investigating intermediate steps in assembly.
Organizational Affiliation:
Department of Molecular Biology and Biochemistry, Wesleyan University, 52 Lawn Avenue, Middletown, CT 06459, USA.