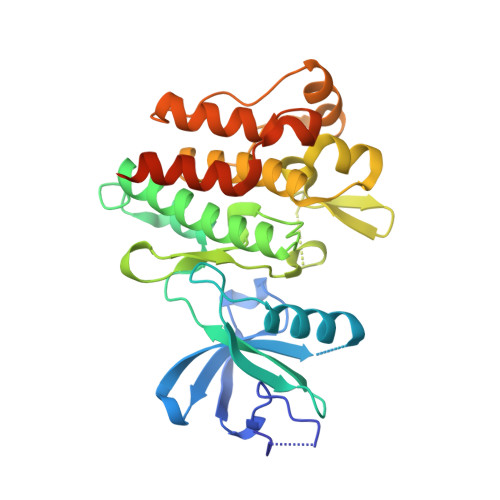
Binding or bending: distinction of allosteric Abl kinase agonists from antagonists by an NMR-based conformational assay.
Jahnke, W., Grotzfeld, R.M., Pelle, X., Strauss, A., Fendrich, G., Cowan-Jacob, S.W., Cotesta, S., Fabbro, D., Furet, P., Mestan, J., Marzinzik, A.L.(2010) J Am Chem Soc 132: 7043-7048
- PubMed: 20450175
- DOI: https://doi.org/10.1021/ja101837n
- Primary Citation of Related Structures:
3MS9, 3MSS - PubMed Abstract:
Allosteric inhibitors of Bcr-Abl have emerged as a novel therapeutic option for the treatment of CML. Using fragment-based screening, a search for novel Abl inhibitors that bind to the myristate pocket was carried out. Here we show that not all myristate ligands are functional inhibitors, but that the conformational state of C-terminal helix_I is a structural determinant for functional activity. We present an NMR-based conformational assay to monitor the conformation of this crucial helix_I and show that myristate ligands that bend helix_I are functional antagonists, whereas ligands that bind to the myristate pocket but do not induce this conformational change are kinase agonists. Activation of c-Abl by allosteric agonists has been confirmed in a biochemical assay.
Organizational Affiliation:
Novartis Institutes for Biomedical Research, 4002 Basel, Switzerland. wolfgang.jahnke@novartis.com