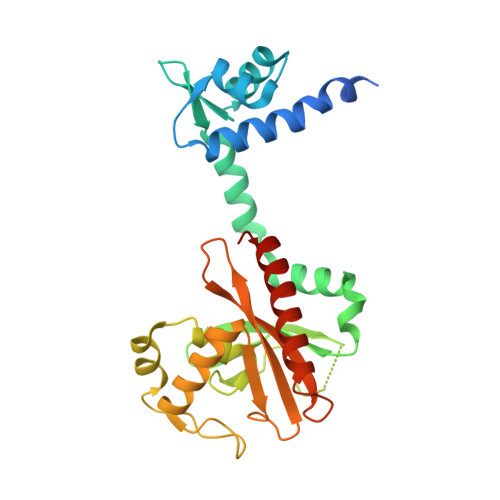
The Agrobacterium tumefaciens Transcription Factor BlcR Is Regulated via Oligomerization.
Pan, Y., Fiscus, V., Meng, W., Zheng, Z., Zhang, L.H., Fuqua, C., Chen, L.(2011) J Biol Chem 286: 20431-20440
- PubMed: 21467043
- DOI: https://doi.org/10.1074/jbc.M110.196154
- Primary Citation of Related Structures:
3MQ0 - PubMed Abstract:
The Agrobacterium tumefaciens BlcR is a member of the emerging isocitrate lyase transcription regulators that negatively regulates metabolism of γ-butyrolactone, and its repressing function is relieved by succinate semialdehyde (SSA). Our crystal structure showed that BlcR folded into the DNA- and SSA-binding domains and dimerized via the DNA-binding domains. Mutational analysis identified residues, including Phe(147), that are important for SSA association; BlcR(F147A) existed as tetramer. Two BlcR dimers bound to target DNA and in a cooperative manner, and the distance between the two BlcR-binding sequences in DNA was critical for BlcR-DNA association. Tetrameric BlcR(F147A) retained DNA binding activity, and importantly, this activity was not affected by the distance separating the BlcR-binding sequences in DNA. SSA did not dissociate tetrameric BlcR(F147A) or BlcR(F147A)-DNA. As well as in the SSA-binding site, Phe(147) is located in a structurally flexible loop that may be involved in BlcR oligomerization. We propose that SSA regulates BlcR DNA-binding function via oligomerization.
Organizational Affiliation:
Indiana University, Bloomington, Indiana 47405, USA.