Selenomethionine incorporation in proteins expressed in Lactococcus lactis.
Berntsson, R.P., Alia Oktaviani, N., Fusetti, F., Thunnissen, A.M., Poolman, B., Slotboom, D.J.(2009) Protein Sci 18: 1121-1127
- PubMed: 19388077
- DOI: https://doi.org/10.1002/pro.97
- Primary Citation of Related Structures:
3FTO - PubMed Abstract:
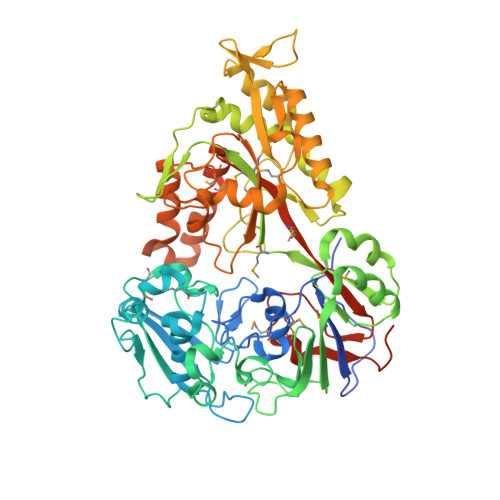
Lactococcus lactis is a promising host for (membrane) protein overproduction. Here, we describe a protocol for incorporation of selenomethionine (SeMet) into proteins expressed in L. lactis. Incorporation efficiencies of SeMet in the membrane protein complex OpuA (an ABC transporter) and the soluble protein OppA, both from L. lactis, were monitored by mass spectrometry. Both proteins incorporated SeMet with high efficiencies (>90%), which greatly extends the usefulness of the expression host L. lactis for X-ray crystallography purposes. The crystal structure of ligand-free OppA was determined at 2.4 A resolution by a semiautomatic approach using selenium single-wavelength anomalous diffraction phasing.
Organizational Affiliation:
Department of Biochemistry, Groningen Biomolecular Sciences and Biotechnology Institute and Zernike Institute for Advanced Materials, University of Groningen, Nijenborgh 4, 9747 AG Groningen, The Netherlands.