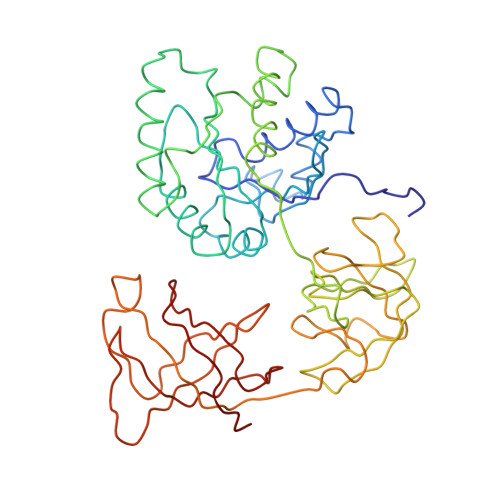
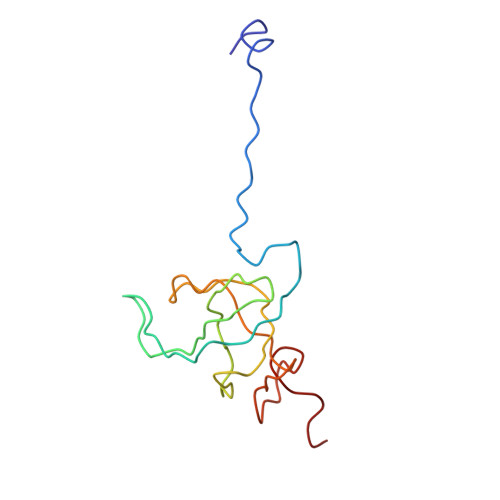
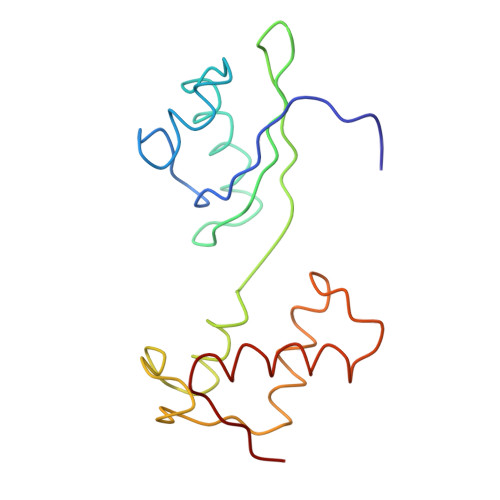
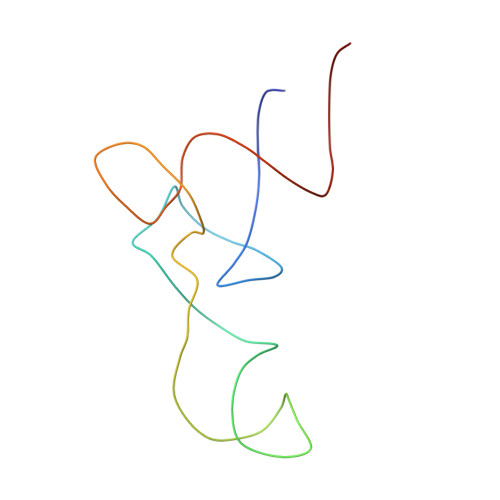Recognition of aminoacyl-tRNA: a common molecular mechanism revealed by cryo-EM.
Li, W., Agirrezabala, X., Lei, J., Bouakaz, L., Brunelle, J.L., Ortiz-Meoz, R.F., Green, R., Sanyal, S., Ehrenberg, M., Frank, J.(2008) EMBO J 27: 3322-3331
- PubMed: 19020518
- DOI: https://doi.org/10.1038/emboj.2008.243
- Primary Citation of Related Structures:
3EP2, 3EQ3, 3EQ4 - PubMed Abstract:
The accuracy of ribosomal translation is achieved by an initial selection and a proofreading step, mediated by EF-Tu, which forms a ternary complex with aminoacyl(aa)-tRNA. To study the binding modes of different aa-tRNAs, we compared cryo-EM maps of the kirromycin-stalled ribosome bound with ternary complexes containing Phe-tRNA(Phe), Trp-tRNA(Trp), or Leu-tRNA(LeuI). The three maps suggest a common binding manner of cognate aa-tRNAs in their specific binding with both the ribosome and EF-Tu. All three aa-tRNAs have the same 'loaded spring' conformation with a kink and twist between the D-stem and anticodon stem. The three complexes are similarly integrated in an interaction network, extending from the anticodon loop through h44 and protein S12 to the EF-Tu-binding CCA end of aa-tRNA, proposed to signal cognate codon-anticodon interaction to the GTPase centre and tune the accuracy of aa-tRNA selection.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA.