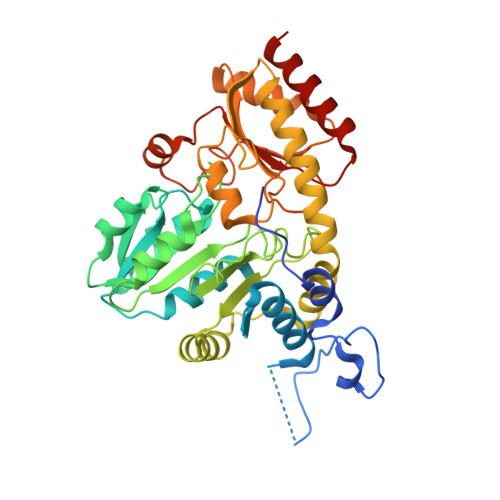
Structural basis for the inhibition mechanism of human cystathionine gamma-lyase, an enzyme responsible for the production of H(2)S.
Sun, Q., Collins, R., Huang, S., Holmberg-Schiavone, L., Anand, G.S., Tan, C.H., van-den-Berg, S., Deng, L.W., Moore, P.K., Karlberg, T., Sivaraman, J.(2009) J Biol Chem 284: 3076-3085
- PubMed: 19019829
- DOI: https://doi.org/10.1074/jbc.M805459200
- Primary Citation of Related Structures:
2NMP, 3COG, 3ELP - PubMed Abstract:
Impairment of the formation or action of hydrogen sulfide (H(2)S), an endogenous gasotransmitter, is associated with various diseases, such as hypertension, diabetes mellitus, septic and hemorrhagic shock, and pancreatitis. Cystathionine beta-synthase and cystathionine gamma-lyase (CSE) are two pyridoxal-5'-phosphate (PLP)-dependent enzymes largely responsible for the production of H(2)S in mammals. Inhibition of CSE by DL-propargylglycine (PAG) has been shown to alleviate disease symptoms. Here we report crystal structures of human CSE (hCSE), in apo form, and in complex with PLP and PLP.PAG. Structural characterization, combined with biophysical and biochemical studies, provides new insights into the inhibition mechanism of hCSE-mediated production of H(2)S. Transition from the open form of apo-hCSE to the closed PLP-bound form reveals large conformational changes hitherto not reported. In addition, PAG binds hCSE via a unique binding mode, not observed in PAG-enzyme complexes previously. The interaction of PAG-hCSE was not predicted based on existing information from known PAG complexes. The structure of hCSE.PLP.PAG complex highlights the particular importance of Tyr(114) in hCSE and the mechanism of PAG-dependent inhibition of hCSE. These results provide significant insights, which will facilitate the structure-based design of novel inhibitors of hCSE to aid in the development of therapies for diseases involving disorders of sulfur metabolism.
Organizational Affiliation:
Department of Biological Sciences, National University of Singapore, 117543 Singapore.