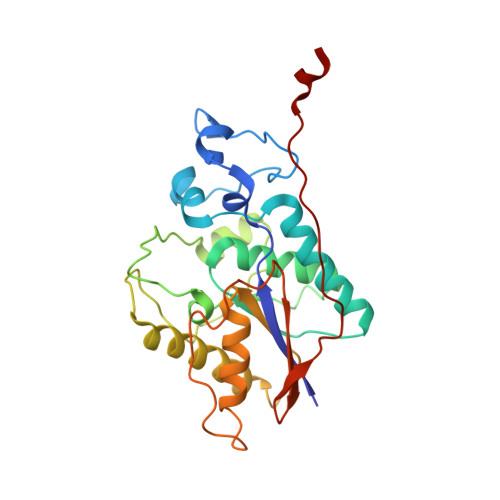
Structural and functional characterization of the c-terminal domain of the ecdysteroid phosphate phosphatase from Bombyx mori reveals a new enzymatic activity.
Chen, Y., Jakoncic, J., Wang, J., Zheng, X., Carpino, N., Nassar, N.(2008) Biochemistry 47: 12135-12145
- PubMed: 18937503
- DOI: https://doi.org/10.1021/bi801318w
- Primary Citation of Related Structures:
3C7T - PubMed Abstract:
Here, we present the crystal structure of the ecdysone phosphate phosphatase (EPPase) phosphoglycerate mutase (PGM) homology domain, the first structure of a steroid phosphate phosphatase. The structure reveals an alpha/beta-fold common to members of the two histidine (2H)-phosphatase superfamily with strong homology to the Suppressor of T-cell receptor signaling-1 (Sts-1 PGM) protein. The putative EPPase PGM active site contains signature residues shared by 2H-phosphatase enzymes, including a conserved histidine (His80) that acts as a nucleophile during catalysis. The physiological substrate ecdysone 22-phosphate was modeled in a hydrophobic cavity close to the phosphate-binding site. EPPase PGM shows limited substrate specificity with an ability to hydrolyze steroid phosphates, the phospho-tyrosine (pTyr) substrate analogue para-nitrophenylphosphate ( pNPP) and pTyr-containing peptides and proteins. Altogether, our data demonstrate a new protein tyrosine phosphatase (PTP) activity for EPPase. They suggest that EPPase and its closest homologues can be grouped into a distinct subfamily in the large 2H-phosphatase superfamily of proteins.
Organizational Affiliation:
Department of Physiology and Biophysics, Stony Brook University, Basic Sciences Tower, Stony Brook, New York 11794-8661, USA.