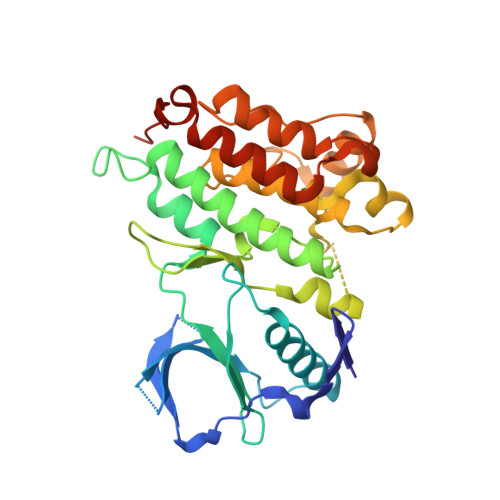
Crystal Structures of Anaplastic Lymphoma Kinase in Complex with ATP Competitive Inhibitors.
Bossi, R.T., Saccardo, M.B., Ardini, E., Menichincheri, M., Rusconi, L., Magnaghi, P., Orsini, P., Avanzi, N., Borgia, A.L., Nesi, M., Bandiera, T., Fogliatto, G., Bertrand, J.A.(2010) Biochemistry 49: 6813
- PubMed: 20695522
- DOI: https://doi.org/10.1021/bi1005514
- Primary Citation of Related Structures:
2XB7, 2XBA - PubMed Abstract:
Anaplastic lymphoma kinase (ALK) is a receptor tyrosine kinase involved in the development of several human cancers and, as a result, is a recognized target for the development of small-molecule inhibitors for the treatment of ALK-positive malignancies. Here, we present the crystal structures of the unphosphorylated human ALK kinase domain in complex with the ATP competitive ligands PHA-E429 and NVP-TAE684. Analysis of these structures provides valuable information concerning the specific characteristics of the ALK active site as well as giving indications about how to obtain selective ALK inhibitors. In addition, the ALK-KD-PHA-E429 structure led to the identification of a potential regulatory mechanism involving a link made between a short helical segment immediately following the DFG motif and an N-terminal two-stranded beta-sheet. Finally, mapping of the activating mutations associated with neuroblastoma onto our structures may explain the roles these residues have in the activation process.
Organizational Affiliation:
Nerviano Medical Sciences S.r.l., Viale Pasteur 10, 20014 Nerviano (MI), Italy.