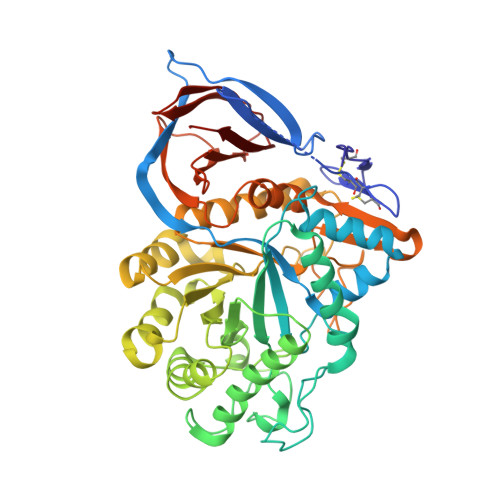
Acid Beta-Glucosidase: Insights from Structural Analysis and Relevance to Gaucher Disease Therapy.
Kacher, Y., Brumshtein, B., Boldin-Adamsky, S., Toker, L., Shainskaya, A., Silman, I., Sussman, J.L., Futerman, A.H.(2008) Biol Chem 389: 1361
- PubMed: 18783340
- DOI: https://doi.org/10.1515/BC.2008.163
- Primary Citation of Related Structures:
2VT0 - PubMed Abstract:
In mammalian cells, glucosylceramide (GlcCer), the simplest glycosphingolipid, is hydrolyzed by the lysosomal enzyme acid beta-glucosidase (GlcCerase). In the human metabolic disorder Gaucher disease, GlcCerase activity is significantly decreased owing to one of approximately 200 mutations in the GlcCerase gene. The most common therapy for Gaucher disease is enzyme replacement therapy (ERT), in which patients are given intravenous injections of recombinant human GlcCerase; the Genzyme product Cerezyme has been used clinically for more than 15 years and is administered to approximately 4000 patients worldwide. Here we review the crystal structure of Cerezyme and other recombinant forms of GlcCerase, as well as of their complexes with covalent and non-covalent inhibitors. We also discuss the stability of Cerezyme, which can be altered by modification of its N-glycan chains with possible implications for improved ERT in Gaucher disease.
Organizational Affiliation:
Department of Biological Chemistry, Weizmann Institute of Science, Rehovot 76100, Israel.