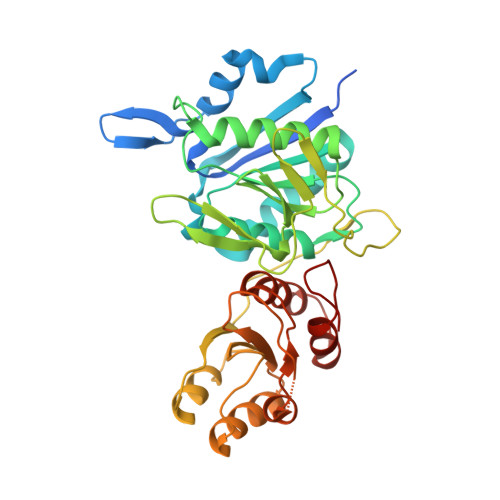
Crystal Structure and Allosteric Regulation of the Cytoplasmic Escherichia colil-Asparaginase I
Yun, M.-K., Nourse, A., White, S.W., Rock, C.O., Heath, R.J.(2007) J Mol Biol 369: 794-811
- PubMed: 17451745
- DOI: https://doi.org/10.1016/j.jmb.2007.03.061
- Primary Citation of Related Structures:
2HIM, 2P2D, 2P2N - PubMed Abstract:
AnsA is the cytoplasmic asparaginase from Escherichia coli involved in intracellular asparagine utilization. Analytical ultracentifugation and X-ray crystallography reveal that AnsA forms a tetrameric structure as a dimer of two intimate dimers. Kinetic analysis of the enzyme reveals that AnsA is positively cooperative, displaying a sigmoidal substrate dependence curve with an [S](0.5) of 1 mM L-asparagine and a Hill coefficient (n(H)) of 2.6. Binding of L-asparagine to an allosteric site was observed in the crystal structure concomitant with a reorganization of the quarternary structure, relative to the apo enzyme. The carboxyl group of the bound asparagine makes salt bridges and hydrogen bonds to Arg240, while the N(delta2) nitrogen interacts with Thr162. Mutation of Arg240 to Ala increases the [S](0.5) value to 5.9 mM, presumably by reducing the affinity of the site for L-asparagine, although the enzyme retains cooperativity. Mutation of Thr162 to Ala results in an active enzyme with no cooperativity. Transmission of the signal from the allosteric site to the active site appears to involve subtle interactions at the dimer-dimer interface and relocation of Gln118 into the vicinity of the active site to position the probable catalytic water molecule. These data define the structural basis for the cooperative regulation of the intracellular asparaginase that is required for proper functioning within the cell.
Organizational Affiliation:
Department of Structural Biology, St Jude Children's Research Hospital, Memphis, TN 38105, USA.