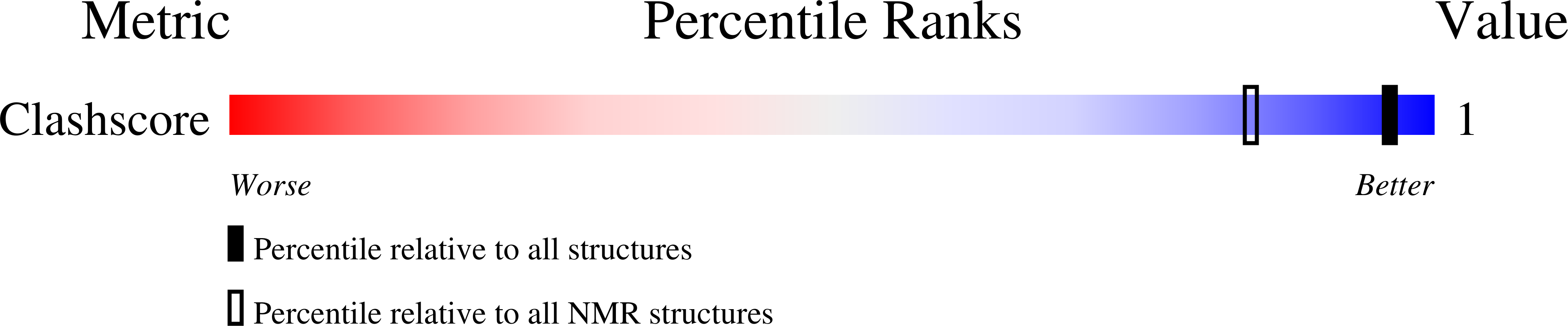A minimal i-motif stabilized by minor groove G:T:G:T tetrads.
Escaja, N., Viladoms, J., Garavis, M., Villasante, A., Pedroso, E., Gonzalez, C.(2012) Nucleic Acids Res 40: 11737-11747
- PubMed: 23042679
- DOI: https://doi.org/10.1093/nar/gks911
- Primary Citation of Related Structures:
2LSX - PubMed Abstract:
The repetitive DNA sequences found at telomeres and centromeres play a crucial role in the structure and function of eukaryotic chromosomes. This role may be related to the tendency observed in many repetitive DNAs to adopt non-canonical structures. Although there is an increasing recognition of the importance of DNA quadruplexes in chromosome biology, the co-existence of different quadruplex-forming elements in the same DNA structure is still a matter of debate. Here we report the structural study of the oligonucleotide d(TCGTTTCGT) and its cyclic analog d
. Both sequences form dimeric quadruplex structures consisting of a minimal i-motif capped, at both ends, by a slipped minor groove-aligned G:T:G:T tetrad. These mini i-motifs, which do not exhibit the characteristic CD spectra of other i-motif structures, can be observed at neutral pH, although they are more stable under acidic conditions. This finding is particularly relevant since these oligonucleotide sequences do not contain contiguous cytosines. Importantly, these structures resemble the loop moiety adopted by an 11-nucleotide fragment of the conserved centromeric protein B (CENP-B) box motif, which is the binding site for the CENP-B.
Organizational Affiliation:
Departament de Química Orgànica and IBUB, Universitat de Barcelona, Martí i Franquès 1-11, 08028 Barcelona, Spain.