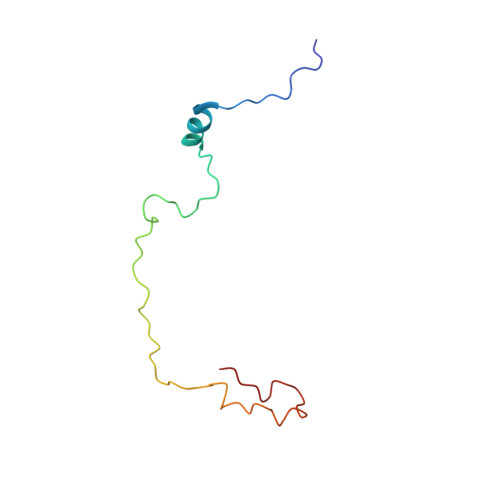
Micelle-induced folding of spinach thylakoid soluble phosphoprotein of 9 kDa and its functional implications.
Song, J., Lee, M.S., Carlberg, I., Vener, A.V., Markley, J.L.(2006) Biochemistry 45: 15633-15643
- PubMed: 17176085
- DOI: https://doi.org/10.1021/bi062148m
- Primary Citation of Related Structures:
2FFT - PubMed Abstract:
Thylakoid soluble phosphoprotein of 9 kDa (TSP9) has been identified as a plant-specific protein in the photosynthetic thylakoid membrane (Carlberg et al. (2003) Proc. Natl. Acad. Sci. 100, 757-762). Nonphosphorylated TSP9 is associated with the membrane, whereas, after light-induced phosphorylation, a fraction of the phosphorylated TSP9 is released into the aqueous stroma. By NMR spectroscopy, we have determined the structural features of nonphosphorylated TSP9 both in aqueous solution and in membrane mimetic micelles. The results show that both wild type nonphosphorylated TSP9 and a triple-mutant (T46E + T53E + T60E) mimic of the triphosphorylated form of TSP9 are disordered under aqueous conditions, but adopt an ordered conformation in the presence of detergent micelles. The micelle-induced structural features, which are similar in micelles either of SDS or dodecylphosphocholine (DPC), consist of an N-terminal alpha-helix, which may represent the primary site of interaction between TSP9 and binding partners, and a less structured helical turn near the C-terminus. These structured elements contain mainly hydrophobic residues. NMR relaxation data for nonphosphorylated TSP9 in SDS micelles indicated that the molecule is highly flexible with the highest order in the N-terminal alpha-helix. Intermolecular NOE signals, as well as spin probe-induced broadening of NMR signals, demonstrated that the SDS micelles contact both the structured and a portion of the unstructured regions of TSP9, in particular, those containing the three phosphorylation sites (T46, T53, and T60). This interaction may explain the selective dissociation of phosphorylated TSP9 from the membrane. Our study presents a structural model for the role played by the structured and unstructured regions of TSP9 in its membrane association and biological function.
Organizational Affiliation:
Center for Eukaryotic Structural Genomics, Department of Biochemistry, University of Wisconsin-Madison, Wisconsin 53706-1544, USA.