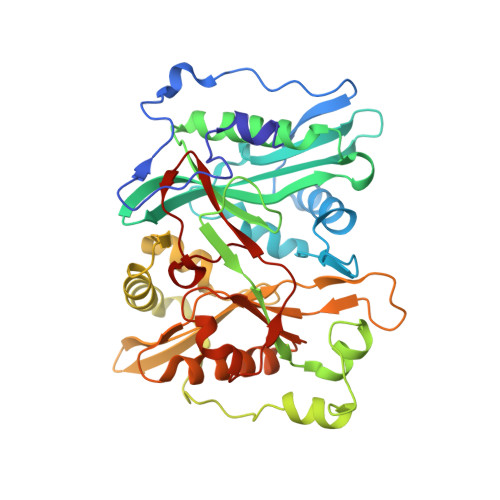
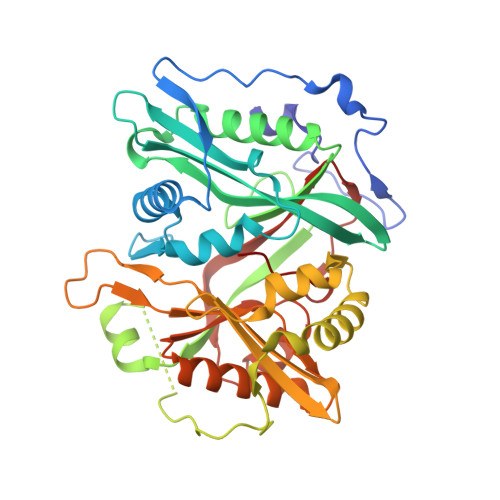
Validation of N-Myristoyltransferase as an Antimalarial Drug Target Using an Integrated Chemical Biology Approach.
Wright, M.H., Clough, B., Rackham, M.D., Rangachari, K., Brannigan, J.A., Grainger, M., Moss, D.K., Bottrill, A.R., Heal, W.P., Broncel, M., Serwa, R.A., Brady, D., Mann, D., Leatherbarrow, R.J., Tewari, R., Wilkinson, A.J., Holder, A.A., Tate, E.W.(2014) Nat Chem 6: 112
- PubMed: 24451586
- DOI: https://doi.org/10.1038/nchem.1830
- Primary Citation of Related Structures:
2YNC, 2YND, 2YNE - PubMed Abstract:
Malaria is an infectious disease caused by parasites of the genus Plasmodium, which leads to approximately one million deaths per annum worldwide. Chemical validation of new antimalarial targets is urgently required in view of rising resistance to current drugs. One such putative target is the enzyme N-myristoyltransferase, which catalyses the attachment of the fatty acid myristate to protein substrates (N-myristoylation). Here, we report an integrated chemical biology approach to explore protein myristoylation in the major human parasite P. falciparum, combining chemical proteomic tools for identification of the myristoylated and glycosylphosphatidylinositol-anchored proteome with selective small-molecule N-myristoyltransferase inhibitors. We demonstrate that N-myristoyltransferase is an essential and chemically tractable target in malaria parasites both in vitro and in vivo, and show that selective inhibition of N-myristoylation leads to catastrophic and irreversible failure to assemble the inner membrane complex, a critical subcellular organelle in the parasite life cycle. Our studies provide the basis for the development of new antimalarials targeting N-myristoyltransferase.
Organizational Affiliation:
1] Department of Chemistry, Imperial College London, London SW7 2AZ, UK [2] Institute of Chemical Biology, Imperial College London, London SW7 2AZ, UK.