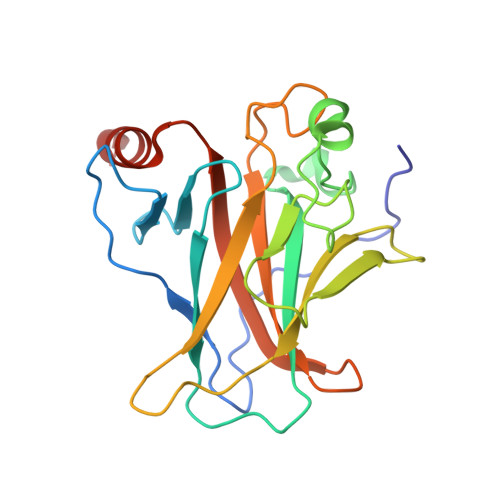
Interaction of the P53 DNA-Binding Domain with its N-Terminal Extension Modulates the Stability of the P53 Tetramer.
Natan, E., Baloglu, C., Pagel, K., Freund, S.M., Morgner, N., Robinson, C.V., Fersht, A.R., Joerger, A.C.(2011) J Mol Biol 409: 358
- PubMed: 21457718
- DOI: https://doi.org/10.1016/j.jmb.2011.03.047
- Primary Citation of Related Structures:
2XWR - PubMed Abstract:
The tetrameric tumor suppressor p53 plays a pivotal role in the control of the cell cycle and provides a paradigm for an emerging class of oligomeric, multidomain proteins with structured and intrinsically disordered regions. Many of its biophysical and functional properties have been extrapolated from truncated variants, yet the exact structural and functional role of certain segments of the protein is unclear. We found from NMR and X-ray crystallography that the DNA-binding domain (DBD) of human p53, usually defined as residues 94-292, extends beyond these domain boundaries. Trp91, in the hinge region between the disordered proline-rich N-terminal domain and the DBD, folds back onto the latter and has a cation-π interaction with Arg174. These additional interactions increase the melting temperature of the DBD by up to 2 °C and inhibit aggregation of the p53 tetramer. They also modulate the dissociation of the p53 tetramer. The absence of the Trp91/Arg174 packing presumably allows nonnative DBD-DBD interactions that both nucleate aggregation and stabilize the interface. These data have important implications for studies of multidomain proteins in general, highlighting the fact that weak ordered-disordered domain interactions can modulate the properties of proteins of complex structure.
Organizational Affiliation:
MRC Laboratory of Molecular Biology, Cambridge, UK.